- Chinese Optics Letters
- Vol. 20, Issue 4, 041101 (2022)
Abstract
1. Introduction
Light field (LF) imaging technology has shown great significance in recent years for its high-temporal-resolution 3D imaging feature through simultaneously capturing the 2D spatial and 2D angular information of light [four-dimensional (4D) LF information][
In recent years, deep learning has been widely studied due to its remarkable ability in information processing and has shown great value in the field of computational imaging[
In this paper, a deep learning-based method of removal of the scattering effect on LF imaging is proposed to deal with the case of scattering imaging in LF imaging applications mentioned above. We term this method deep learning-based scattering LF imaging (DeepSLFI). In DeepSLFI, we build the LF imaging forward models and generate training samples of LF images with and without scattering by simulation. By the means of simulation, the huge experimental workload for samples capturing can be avoided. Then, a neural network trained with the simulation samples is utilized to remove the effect of scattering on the LF image captured experimentally. Finally, the high-resolution and high-contrast 3D reconstruction can be conducted with the processed LF image. In this paper, a validation experiment is conducted, and the 3D reconstructions of the scene of imaging through a single scattering layer are obtained. Compared to the results of conventional methods, the 3D reconstruction using DeepSLFI has a higher quality in terms of resolution and contrast.
Sign up for Chinese Optics Letters TOC. Get the latest issue of Chinese Optics Letters delivered right to you!Sign up now
2. Method
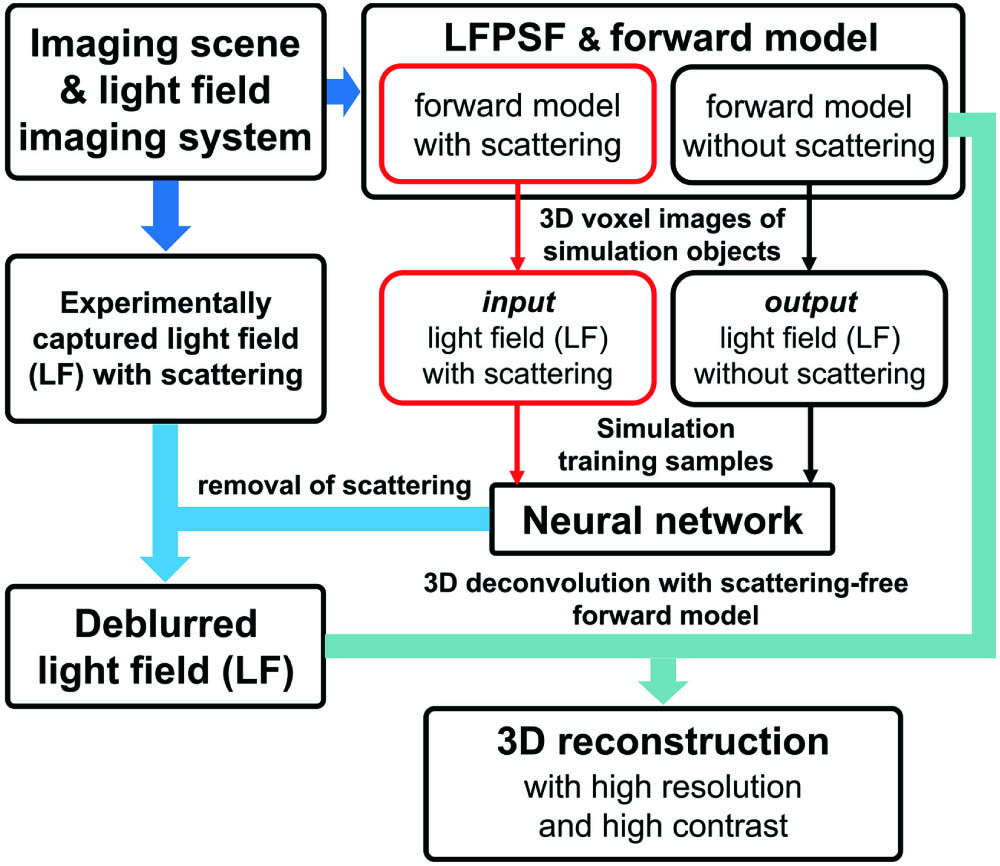
The overview of DeepSLFI is shown in Fig. 1. The building of the LF imaging forward model is conducted with the theory of LF imaging[
![]()
Figure 1.Overview of DeepSLFI. The light field imaging forward models can be built after the scattering imaging scene and the light field imaging system are determined. The simulation light field images serving as training samples can be generated with the forward models. Then, a neural network will be trained with the samples and utilized to remove the scattering of the light field image captured experimentally. Finally, the high-resolution and high-contrast 3D reconstruction can be obtained by 3D deconvolution with the deblurred light field image and the scattering-free forward model.
![]()
Figure 2.Diagram of light field imaging forward model. Light emitted from voxels of object space
In the absence of scattering, light propagation can be described by theories of wave optics and Fresnel propagation[
In the presence of scattering, light propagation can be described by an analysis method based on phase-space theory[
After determining the light propagation in the imaging scene, the intensity proportion on the sensor of the light emitted from point emitter in the object space,
![]()
Figure 3.Light field images captured experimentally of the same object (a) without scattering and (b) with scattering. (a1) and (b1) correspond to the parts of the dashed boxes in (a) and (b), respectively. (c) The intensity curves of pixels on the blue and green lines.
In order to remove the effect of scattering on the LF image, a convolutional neural network (CNN) is used in DeepSLFI. Figure 4 is the overall architecture of the CNN, which is adapted from the residual network (ResNet)[
![]()
Figure 4.Architecture of the neural network.
Considering the huge workload and the serious difficulty for the experimental capturing of samples of 3D images or LF images in scattering scenes, the training sample pairs will be generated by simulation with the LF imaging forward models. This method also makes DeepSLFI more flexible. The LF imaging forward models have been applied in research[
The final reconstruction will be realized by 3D deconvolution using the scattering-removed LF image and the scattering-free forward model (which has sharp patterns of LFPSFs). In this paper, CNN is utilized to remove the scattering of the LF image, not to produce 3D image results of the inverse problem end-to-end[
3. Experiment and Results
An experiment is conducted to verify the validity of DeepSLFI. Compared with the 3D reconstruction results obtained from different LF imaging methods, the advantages of DeepSLFI in LF imaging through scattering could be demonstrated.
The experimental system is shown in Fig. 5, which consists of a CCD sensor (BFLY-PGE-50H5M-C, Point Grey, 3.45 µm pixel size and
![]()
Figure 5.Diagram of the validation experimental system.
The effect of the scattering layer on the LF image is shown in Fig. 3. Considering that the intensity curves in Fig. 3(c) are smooth in general, and the homogeneous scattering approximation in research of LF imaging through scattering has demonstrated its validity[
By calculating the light intensity distributions on the sensor after the propagation shown in Fig. 6, the LFPSFs and the forward models can be obtained with Eqs. (1)–(10). A set of parameters of the scattering model, which can well describe the scattering effect of the scattering layer in the experiment, is used. Generally, the effect of aberration can be negligible in the forward model[
![]()
Figure 6.Light propagation in the experimental system.
The simulation training samples are generated based on the scene: single-layer planar objects located on 25 depths (1 voxel thickness) with intensity maps of the Modified National Institute of Standards and Technology (MNIST)[
![]()
Figure 7.Generation process of the training samples.
Before the neural network training process, all of the generated LF images are scaled to [−1,1] (the range of tanh activation function). Each convolution layer of the network has a stride of 1 and kernel size of
LF images of the object are captured experimentally with and without scattering, respectively. The experiment is conducted under the same condition except for the presence or absence of the scattering layer. The 3D deconvolutions (using the Richardson–Lucy iteration scheme) of the same object are performed in four ways: using the LF without scattering and the scattering-free forward model (recorded as
Figure 8 shows the results of a United States Air Force (USAF) 1951 target (Group 0, Element 6, attached to a white light source to get even illumination) located in the depth of field with a distance of 10 mm from the front plane of the depth of field. To quantitatively analyze the reconstruction results, the peak signal to noise ratio (PSNR) and structural similarity (SSIM)[
![]()
Figure 8.Reconstruction results of the USAF target in the field of view. All of the images are scaled to [0,1]. (a) (i) Light field without scattering, (ii) with scattering, and (iii) deblurred by the network. (iv) The intensity curves of pixels corresponding to the lines in (i), (ii), and (iii). (v) The PSNRs and SSIMs of the LFs. (b) 3D reconstructions from perspective and orthogonal views (removed noise containing only several voxels that interferes with the observation), where the x and y directions are transverse directions, and the z direction is the depth/axial direction. The yellow dashed box shows the depth position of the object. (c) Slice images in the 3D reconstructions. (d) The intensity curves of voxels/pixels corresponding to the dashed lines in the slice images in (c), with (i) the curves of the manganese-purple dashed line in the x-y section, (ii) the curves of the green dashed line in the x-y section, (iii) the curves of the green dashed line in the x-z section, and also the manganese-purple dashed line in the y-z section. (iv) The PSNRs and SSIMs of the 3D reconstructions, where only the part consisting of areas in the yellow dashed box of each depth is selected for calculation. This is because the part outside is the edge of the field of view and has a low quality of reconstruction. It is not suitable for comparison of PSNRs and SSIMs affected by scattering.
Figure 9 shows the 3D reconstruction results of two slits localized at different depths in object space. It shows the ability of 3D reconstruction through scattering of DeepSLFI.
![]()
Figure 9.Results of reconstructions with different ways for two slits to be localized at different depths in object space. (a) The 3D reconstructions. The blue dashed boxes show the sizes and the positions of the slits. The “Depth” means the distance from the front plane of the depth of field. The median filtering is conducted at the edge of the lateral field of view to remove noise containing several voxels. (b) The PSNRs and SSIMs of the 3D reconstructions.
4. Conclusion
In this paper, we propose DeepSLFI to remove the effect of scattering on LF imaging. DeepSLFI utilizes a CNN, trained by simulation samples, which are generated by the LF imaging forward models, to remove the effect of scattering on the LF image captured experimentally and then realizes high-quality 3D reconstruction using the scattering-removed LF image and the scattering-free forward model. The advantage of DeepSLFI in high-quality imaging in terms of resolution and contrast is demonstrated experimentally. The future research of DeepSLFI will focus on the applications with more different and complex imaging scenes, such as scenes with volumetric scattering and stronger scattering. Future research will also focus on specific applications, such as 3D imaging in biological tissue and fog.
References
[1] M. Martínez-Corral, B. Javidi. Fundamentals of 3D imaging and displays: a tutorial on integral imaging, light-field, and plenoptic systems. Adv. Opt. Photon., 10, 512(2018).
[2] G. Lippmann. Epreuves reversibles, photographies integrales. Comptes Rendus l’Académie des Sci., 444, 446(1908).
[3] R. Ng, M. Levoy, M. Brédif, G. Duval, M. Horowitz, P. Hanrahan. Light field photography with a hand-held plenoptic camera(2005).
[4] M. Levoy, R. Ng, A. Adams, M. Footer, M. Horowitz. Light field microscopy. ACM Trans. Graph., 25, 924(2006).
[5] M. Levoy, Z. Zhang, I. McDowall. Recording and controlling the 4D light field in a microscope using microlens arrays. J. Microsc., 235, 144(2009).
[6] M. Broxton, L. Grosenick, S. Yang, N. Cohen, A. Andalman, K. Deisseroth, M. Levoy. Wave optics theory and 3-D deconvolution for the light field microscope. Opt. Express, 21, 25418(2013).
[7] R. Prevedel, Y.-G. Yoon, M. Hoffmann, N. Pak, G. Wetzstein, S. Kato, T. Schrödel, R. Raskar, M. Zimmer, E. S. Boyden, A. Vaziri. Simultaneous whole-animal 3D imaging of neuronal activity using light-field microscopy. Nat. Methods, 11, 727(2014).
[8] H. Y. Liu, E. Jonas, L. Tian, J. Zhong, B. Recht, L. Waller. 3D imaging in volumetric scattering media using phase-space measurements. Opt. Express, 23, 14461(2015).
[9] N. C. Pégard, H. Y. Liu, N. Antipa, M. Gerlock, H. Adesnik, L. Waller. Compressive light-field microscopy for 3D neural activity recording. Optica, 3, 517(2016).
[10] T. Nöbauer, O. Skocek, A. J. Pernía-Andrade, L. Weilguny, F. M. Traub, M. I. Molodtsov, A. Vaziri. Video rate volumetric Ca2+ imaging across cortex using seeded iterative demixing (SID) microscopy. Nat. Methods, 14, 811(2017).
[11] M. A. Taylor, T. Nöbauer, A. Pernia-Andrade, F. Schlumm, A. Vaziri. Brain-wide 3D light-field imaging of neuronal activity with speckle-enhanced resolution. Optica, 5, 345(2018).
[12] Y. Chen, B. Xiong, Y. Xue, X. Jin, J. Greene, L. Tian. Design of a high-resolution light field miniscope for volumetric imaging in scattering tissue. Biomed. Opt. Express, 11, 1662(2020).
[13] Y. Xue, I. Davison, D. Boas, L. Tian. Single-shot 3D wide-field fluorescence imaging with a computational miniature mesoscope. Sci. Adv., 6, eabb7508(2020).
[14] Y. Zhang, Z. Lu, J. Wu, X. Lin, D. Jiang, Y. Cai, J. Xie, Y. Wang, T. Zhu, X. Ji, Q. Dai. Computational optical sectioning with an incoherent multiscale scattering model for light-field microscopy. Nat. Commun., 12, 6391(2021).
[15] I. Moon, B. Javidi. Three-dimensional visualization of objects in scattering medium by use of computational integral imaging. Opt. Express, 16, 13080(2008).
[16] J. Tian, Z. Murez, T. Cui, Z. Zhang, D. Kriegman, R. Ramamoorthi. Depth and image restoration from light field in a scattering medium. IEEE Conference ICCV, 2420(2017).
[17] K. Yanny, N. Antipa, W. Liberti, S. Dehaeck, K. Monakhova, F. L. Liu, K. Shen, R. Ng, L. Waller. Miniscope3D: optimized single-shot miniature 3D fluorescence microscopy. Light Sci. Appl., 9, 171(2020).
[18] G. Barbastathis, A. Ozcan, G. Situ. On the use of deep learning for computational imaging. Optica, 6, 921(2019).
[19] Y. Rivenson, Z. Gorocs, H. Gunaydin, Y. Zhang, H. Wang, A. Ozcan. Deep learning microscopy. Optica, 4, 1437(2017).
[20] E. Nehme, L. E. Weiss, T. Michaeli, Y. Shechtman. Deep-STORM: super-resolution single-molecule microscopy by deep learning. Optica, 5, 458(2018).
[21] A. Sinha, J. Lee, S. Li, G. Barbastathis. Lensless computational imaging through deep learning. Optica, 4, 1117(2017).
[22] F. Wang, H. Wang, H. Wang, G. Li, G. Situ. Learning from simulation: an end-to-end deep-learning approach for computational ghost imaging. Opt. Express, 27, 25560(2019).
[23] F. Wang, Y. Bian, H. Wang, M. Lyu, G. Pedrini, W. Osten, G. Barbastathis, G. Situ. Phase imaging with an untrained neural network. Light Sci. Appl., 9, 77(2020).
[24] S. Yuan, Y. Hu, Q. Hao, S. Zhang. High-accuracy phase demodulation method compatible to closed fringes in a single-frame interferogram based on deep learning. Opt. Express, 29, 2538(2021).
[25] Z. Wang, L. Zhu, H. Zhang, G. Li, C. Yi, Y. Li, Y. Yang, Y. Ding, M. Zhen, S. Gao, T. K. Hsiai, P. Fei. Real-time volumetric reconstruction of biological dynamics with light-field microscopy and deep learning. Nat. Methods, 18, 551(2021).
[26] N. Wagner, F. Beuttenmueller, N. Norlin, J. Gierten, J. C. Boffi, J. Wittbrodt, M. Weigert, L. Hufnagel, R. Prevedel, A. Kreshuk. Deep learning-enhanced light-field imaging with continuous validation. Nat. Methods, 18, 557(2021).
[27] S. Li, M. Deng, J. Lee, A. Sinha, G. Barbastathis. Imaging through glass diffusers using densely connected convolutional networks. Optica, 5, 803(2018).
[28] Y. Li, Y. Xue, L. Tian. Deep speckle correlation: a deep learning approach toward scalable imaging through scattering media. Optica, 5, 1181(2018).
[29] M. Lyu, H. Wang, G. Li, S. Zheng, G. Situ. Learning-based lensless imaging through optically thick scattering media. Adv. Photon., 1, 036002(2019).
[30] X. Lai, Q. Li, Z. Chen, X. Shao, J. Pu. Reconstructing images of two adjacent objects passing through scattering medium via deep learning. Opt. Express, 29, 43280(2021).
[31] H. Li, C. Guo, D. Kim-Holzapfel, W. Li, Y. Altshuller, B. Schroeder, W. Liu, Y. Meng, J. B. French, K.-I. Takamaru, M. A. Frohman, S. Jia. Fast, volumetric live-cell imaging using high-resolution light-field microscopy. Biomed. Opt. Express, 10, 29(2019).
[32] M. J. Bastiaans. The Wigner distribution function applied to optical signals and systems. Opt. Commun., 25, 26(1978).
[33] J. Jönsson, E. Berrocal. Multi-scattering software: part I: online accelerated Monte Carlo simulation of light transport through scattering media. Opt. Express, 28, 37612(2020).
[34] K. He, X. Zhang, S. Ren, J. Sun. Deep residual learning for image recognition. IEEE Conference on Computer Vision and Pattern Recognition, 770(2016).
[35] O. Ronneberger, P. Fischer, T. Brox. U-net: convolutional networks for biomedical image segmentation. International Conference on Medical Image Computing and Computer-Assisted Intervention, 234(2015).
[36] W. Wang, Z. Wang, Y. Wen, L. Song, X. Zhao, J. Yang. A method of 3D light field imaging through single layer of weak scattering media basd on deep learning. Proc. SPIE, 11438, 114380Y(2020).
[38] A. G. Asuero, A. Sayago, A. G. González. The correlation coefficient: an overview. Crit. Rev. Anal. Chem., 36, 41(2006).
[39] W. Zhou, A. C. Bovik. Mean squared error: love it or leave it? A new look at Signal fidelity measures. IEEE Signal Process. Mag., 26, 98(2009).
Set citation alerts for the article
Please enter your email address



