Jinge Yang, Seongwook Choi, Jiwoong Kim, Byullee Park, Chulhong Kim. Recent advances in deep-learning-enhanced photoacoustic imaging[J]. Advanced Photonics Nexus, 2023, 2(5): 054001
Search by keywords or author
- Advanced Photonics Nexus
- Vol. 2, Issue 5, 054001 (2023)
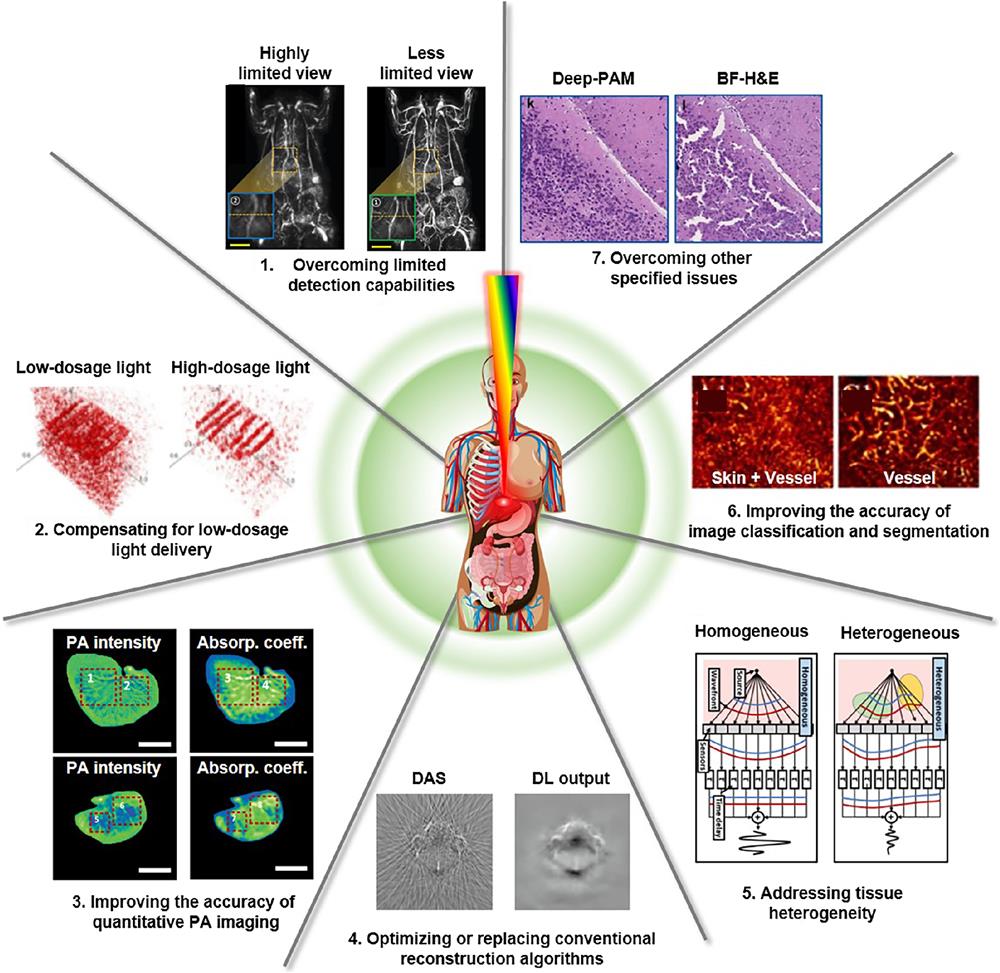
Fig. 1. Representations of seven major challenges in PAI, and DL-related methods to overcome them. DAS, delay-and-sum; DL, deep learning; BF-H&E, bright-field hematoxylin and eosin staining. The images are adapted with permission from Ref. 52, © 2021 Wiley-VCH GmbH; Ref. 53, © 2020 Optica; Ref. 54, © 2022 Optica; Ref. 55, © 2020 Elsevier GmbH; Ref. 56, CC-BY; Ref. 57, © 2021 Elsevier GmbH; and Ref. 58, © 2021 Elsevier GmbH.
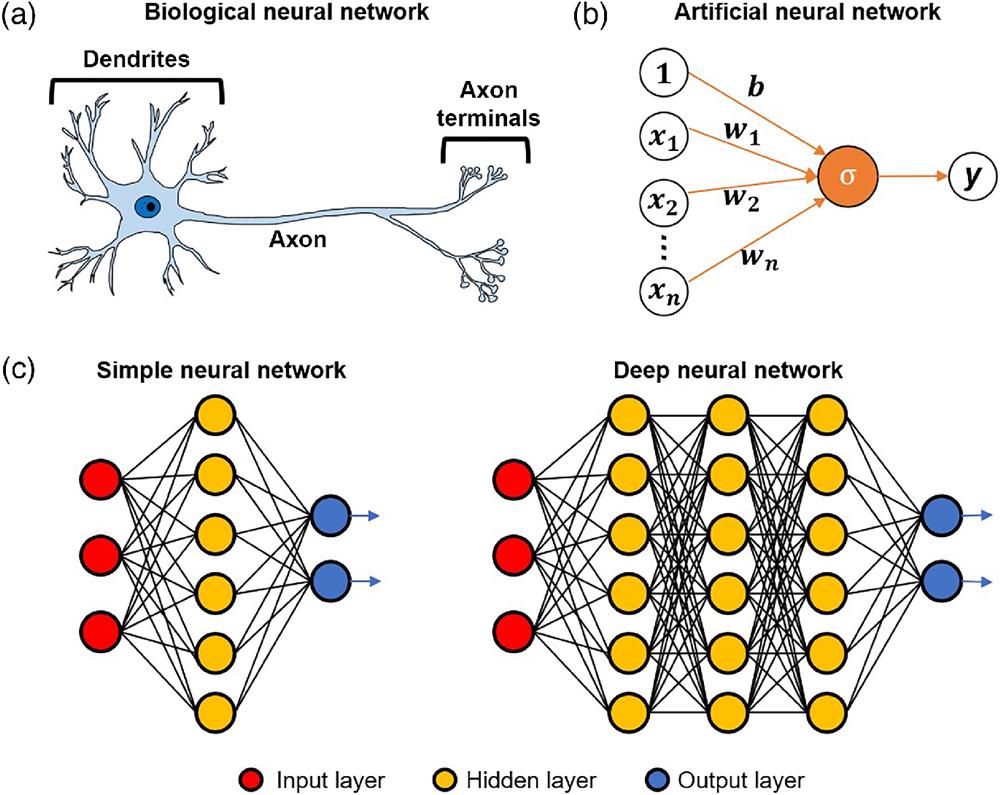
Fig. 2. The concept of (a) a biological neural network and (b) an ANN derived from (a). (c) Schematics of a simple neural network and a DNN.
Fig. 3. Three typical neural network architectures for biomedical imaging. (a) CNN, (b) U-Net, and (c) GAN.
Fig. 4. Representative studies using DL methods to overcome limited-detection capabilities. (a) A DNN with five fully connected layers enhances bandwidth. (b) LV-GAN for addressing the limited-view problem. (c) A Y-Net generates the PA images by optimizing both raw data and reconstructed images from the traditional method. (d) A 3D progressive U-Net (3D-pUnet) to diminish the effects of limited-view artifacts and sparsity arising from cluster view detection. The images are adapted with permission from Ref. 76, © 2017 SPIE; Ref. 77, © 2020 Wiley-VCH GmbH; Ref. 78, © 2020 Elsevier GmbH; and Ref. 52, © 2021 Wiley-VCH GmbH. BW, bandwidth; DNN, deep neural network; DAS, delay-and-sum; cluster, cluster view detection; full, full view detection.
Fig. 5. Representative DL approaches compensate for low laser dosage. (a) An MWCNN that generates high-quality PA images from low-fluence PA images. (b) An HD-UNet that enhances the image quality in a pulsed-laser diode PACT system. (c) An MT-RDN that performs image denoising, superresolution, and vascular enhancement. The images are adapted with permission from Ref. 53, © 2020 Optica; Ref. 109, © 2022 SPIE; and Ref. 110, © 2020 Wiley-VCH GmbH. MWCNN, multi-level wavelet-convolutional neural network; HD-UNet, hybrid dense U-Net; MT-RDN, multitask residual dense network.
Fig. 6. Representative studies to improve the accuracy of quantitative PAI by DL. (a) Convolutional encoder–decoder type network with skip connections (EDS) to produce accurate estimates of
Fig. 7. Representative studies to optimize conventional reconstruction algorithms or replace them with DL. (a) Pixel-wise interpolation approach followed by an FD-UNet for limited-view and sparse PAT image reconstruction. (b) End-to-end U-Net with residual blocks to reconstruct PA images. (c) Two-step PA image reconstruction process with FPnet and U-Net. The images are adapted with permission from Ref. 137, © 2020 Nature Publishing Group; Ref. 138, © 2020 Optica; and Ref. 55, © 2020 Elsevier GmbH.
Fig. 8. Representative DL studies to correct the SoS and improve the accuracy of image classification and segmentation. (a) Hybrid DNN model including U-Net and Segnet to mitigate SOS aberration in heterogeneous tissue. (b) Sparse-UNet (S-UNet) for automatic vascular segmentation in MSOT images. The images are adapted with permission from Ref. 153, CC-BY; Ref. 154, © Elsevier GmbH.
Fig. 9. Representative studies using DL to solve specific issues. (a) GAN-based framework (Wasserstein GAN) to enhance the spatial resolution of AR-PAM. (b) GAN with U-Net to reconstruct superresolution images from raw image frames. (c) Deep-PAM generates virtually stained histological images for both thin sections and thick fresh tissue specimens. The images are adapted with permission from Ref. 178, © 2021 Elsevier GmbH; Ref. 180, © 2022 Springer Nature; and Ref. 58, © 2021 Elsevier GmbH. BF-H&E, brightfield hematoxylin and eosin staining; DNN, deep neural network.
|
Table 1. Summary of challenges facing PAI.
|
Table 2. Three representative networks.
|
Table 3. Summary of overcoming the limited detection capabilities with DL approaches.
|
Table 4. Summary of studies on compensating for low-dosage light delivery.
|
Table 5. Summary of studies to improve the accuracy of quantitative PAI.
|
Table 6. Summary of methods to optimize or replace conventional reconstruction algorithms.
|
Table 7. Summary of methods for addressing tissue heterogeneity.
|
Table 8. Summary of methods for improving the accuracy of image classification and segmentation.
|
Table 9. Summary of methods for addressing other specified issues.

Set citation alerts for the article
Please enter your email address



