Photoacoustic imaging (PAI), recognized as a promising biomedical imaging modality for preclinical and clinical studies, uniquely combines the advantages of optical and ultrasound imaging. Despite PAI’s great potential to provide valuable biological information, its wide application has been hindered by technical limitations, such as hardware restrictions or lack of the biometric information required for image reconstruction. We first analyze the limitations of PAI and categorize them by seven key challenges: limited detection, low-dosage light delivery, inaccurate quantification, limited numerical reconstruction, tissue heterogeneity, imperfect image segmentation/classification, and others. Then, because deep learning (DL) has increasingly demonstrated its ability to overcome the physical limitations of imaging modalities, we review DL studies from the past five years that address each of the seven challenges in PAI. Finally, we discuss the promise of future research directions in DL-enhanced PAI.
- Advanced Photonics Nexus
- Vol. 2, Issue 5, 054001 (2023)
Abstract
Author Presentation Playback
1 Introduction
Photoacoustic imaging (PAI) is a noninvasive and radiation-free biomedical imaging modality that provides high spatial resolution, deep penetration, and great optical absorption contrast by synergistically combining optics and acoustics.1 PAI is based on the photoacoustic (PA) effect, in which optical energy from a pulse laser is converted into acoustic energy waves by the light absorption characteristics of biomolecules.2 The initial pressure of a generated PA wave can be calculated as
Photoacoustic microscopy (PAM) and photoacoustic computed tomography (PACT) are the main PAI modalities. PAM is subdivided into two types, depending on which of the two co-aligned acoustic and optical components is more tightly focused.31
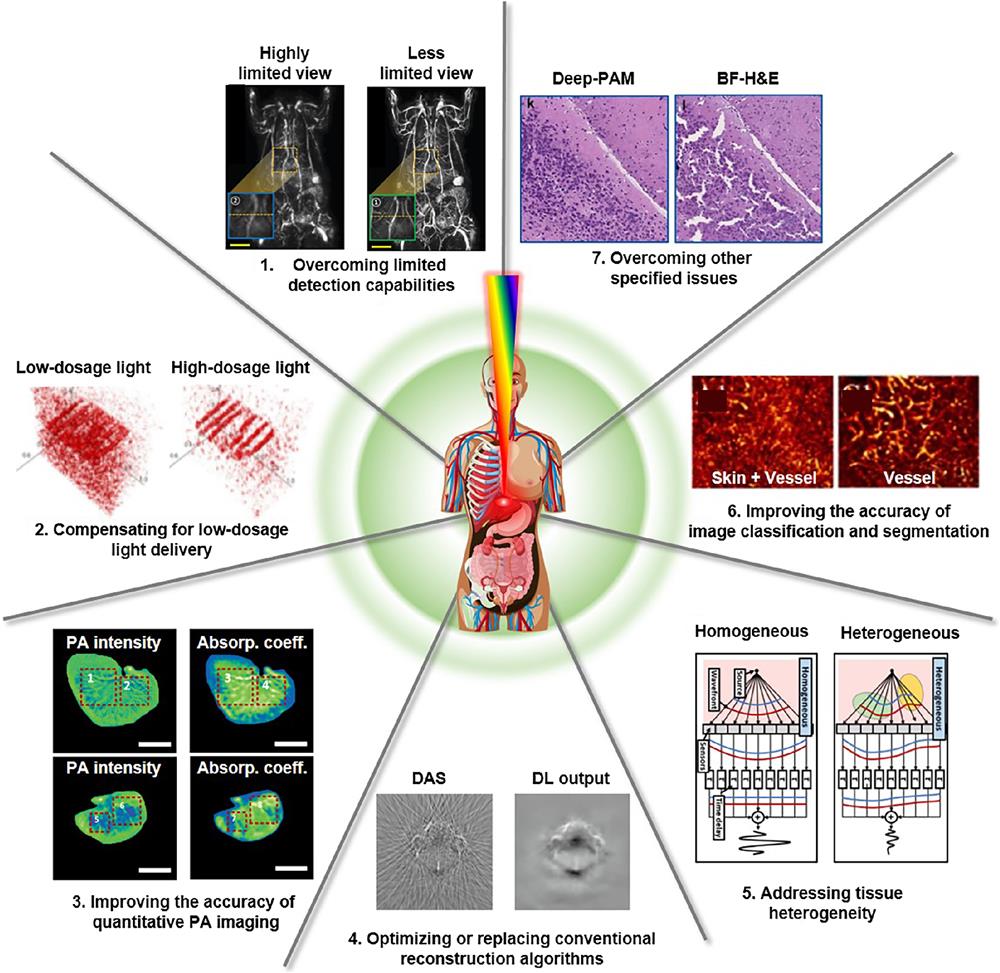
PAI has gained widespread recognition as a promising biomedical imaging modality for preclinical and clinical studies. However, to fully realize PAI’s great potential, the seven challenges listed in Table 1 and illustrated in Fig. 1 must be addressed to further enhance the image quality and expand PAI’s applications:
Sign up for Advanced Photonics Nexus TOC. Get the latest issue of Advanced Photonics Nexus delivered right to you!Sign up now
| Section | Title | Challenges to be solved |
| 3.1 | Overcoming limited detection capabilities | Restricted bandwidth, limited detection view, sampling sparsity |
| 3.2 | Compensating for low-dosage light delivery | Low SNR in the low-dosage light-delivery system |
| 3.3 | Improving the accuracy of quantitative PA imaging | Inaccuracy in quantitative estimates ( |
| 3.4 | Optimizing or replacing conventional reconstruction algorithms | Limitations in conventional reconstruction algorithms |
| 3.5 | Addressing tissue heterogeneity | Acoustic reflection and imaging artifacts led by tissue heterogeneity |
| 3.6 | Improving the accuracy of image classification and segmentation | Inaccuracy and rough classification and segmentation of PA image |
| 3.7 | Overcoming other specified issues | Motion artifacts, limited spatial resolution, electrical noise and interference, image misalignment, accelerating superresolution imaging, achieving digital histologic staining |
Table 1. Summary of challenges facing PAI.
![]()
Figure 1.Representations of seven major challenges in PAI, and DL-related methods to overcome them. DAS, delay-and-sum; DL, deep learning; BF-H&E, bright-field hematoxylin and eosin staining. The images are adapted with permission from Ref.
Overcoming these challenges is important because relying solely on hardware improvements will not be enough to resolve them. It will require significant investments of time and resources to find effective solutions. Deep learning (DL) plays a crucial role in advancing the field of medical and bioimaging by not only addressing the inherent limitations of imaging systems but also by driving substantial improvements in classification and segmentation performance. In recent years, DL has gained significant traction in PAI research, leading to remarkable breakthroughs and achievements. This comprehensive review article provides an in-depth analysis of diverse methodologies and outcomes showcasing the utilization of DL techniques to effectively address the seven challenges encountered in PAI, as previously outlined.
2 Principles of DL Methods
DL is a subset of machine-learning algorithms that encompasses supervised learning, unsupervised learning, and reinforcement learning. Supervised learning is a modeling technique that establishes a correlation between input data and their corresponding ground truth (GT). This approach is commonly utilized in DL-enhanced medical imaging, where high- and low-quality images can be paired. On the other hand, unsupervised learning identifies specific patterns hidden within data, without the use of labeled examples or a priori known answers. Lastly, in reinforcement learning, an algorithm maximizes the final reward by learning through rewards obtained as a result of performing specific actions in a particular environment. A notable example of such a learning algorithm is AlphaGo, the first computer program to beat a human champion Go player.63 Next, we explain the basic structure of DL network and the basic operating principle of DL training. In addition, we introduce the representative DL architectures that are most widely used in the field of image and video: convolution neural network (CNN), U-shaped neural network (U-Net), and generative adversarial network (GAN) architecture.
2.1 Artificial Neural Networks
Artificial neural networks (ANNs) draw inspiration from biological neural networks, wherein different stimuli enter neurons through dendrites and are transmitted to other cells through axons once a threshold of activation is achieved [Fig. 2(a)]. In ANNs, an artificial neuron is a mathematical function conceived as a biological neuron. This function multiplies the various inputs by each weight, sums them, and adds the deviation.
![]()
Figure 2.The concept of (a) a biological neural network and (b) an ANN derived from (a). (c) Schematics of a simple neural network and a DNN.
This sum is then sent to a specific activation function and produces an output [Fig. 2(b)]. This artificial neuron is expressed as
ANNs typically have an input layer, a hidden layer, and an output layer, each comprising multiple units. The number of hidden layers between the input and output determines whether the ANN is a simple neural network or a deep neural network (DNN) [Fig. 2(c)]. Formulaically, the two networks in Fig. 2(c) are described as
2.2 Backpropagation
Backpropagation serves as the fundamental principle for training DL models. In order to grasp the concept of backpropagation, it is necessary to first understand forward propagation. Forward propagation involves sequentially passing an input value through multiple hidden layers to generate an output. For instance, given an input
The learning rate, a hyperparameter that determines the variable’s update amount in proportion to the calculated slope, is set before the learning process and remains unchanged. The number of hidden layers and their dimensions are also hyperparameters. In the following section, we introduce representative ANN architectures commonly used in biomedical imaging, including PAI.
2.3 CNN
CNNs, the most basic DL architecture, have received significant attention in the field of PAI due to their extensive use in image processing and computer vision. CNNs were developed to extract features or patterns in local areas of an image. A convolution operation is a mathematical process that measures the similarity between two functions. The convolution operation in image processing is the process of calculating how well a subsection of an image matches a filter (also referred to as a kernel) and is used for things such as edge filtering. In CNN, the network is trained by learning this filter, which is used to extract image features, as a weight. The encoder–decoder CNN architecture is a relatively simple network structure capable of performing image-to-image translation tasks [Fig. 3(a)].68 Initially, the input image undergoes a series of downsampling and convolution operations. Throughout this process, the dimensions of the image progressively decrease while the number of image channels, representing an additional dimension, increases. This results in a bottleneck in the representation, which is subsequently reversed through a sequence of upsampling and convolutional operations. The bottleneck enforces the network to encode the image into a compact set of abstracted variables (also referred to as latent variables) along the channel dimension. The predicted image is synthesized by decoding these variables in the second half of the network.
![]()
Figure 3.Three typical neural network architectures for biomedical imaging. (a) CNN, (b) U-Net, and (c) GAN.
2.4 U-Net
A U-Net [Fig. 3(b)] is a CNN-based model that was originally proposed for image segmentation in the biomedical field.69 The U-Net is composed of two symmetric networks: a network for obtaining overall context information of an image and a second network for accurate localization. The left part of the U-Net is the encoding process, which encodes the input image to obtain overall context information. The right part of U-Net is the decoding process, which decodes the encoded context information to generate a segmented image. The feature maps obtained during the encoding process are concatenated with up-convolved feature maps at each expanding step in the decoding process, using skip connections.70 This enables the decoder to make more accurate predictions by directly conveying important information in the image. As a result, the U-Net architecture has shown excellent performance in several biomedical image segmentation tasks, even when trained on a very small amount of data, due to data augmentation techniques.
2.5 GAN
A GAN is a type of generative model that learns data through a competition between a generator network and a discriminator network [Fig. 3(c)].71,72 The generator network generates the fake data, and the discriminator network tries to distinguish the real data from the fake data. To deceive the discriminator, the generator aims to generate data that look as realistic as possible, while the discriminator attempts to distinguish the real data from the realistic fake data. Through this competition, both networks learn and improve iteratively, resulting in a generator that can generate increasingly realistic data. As a result, GANs have been successful in generating synthetic data that are very similar to real data, making them useful for applications such as data augmentation and image synthesis. These three representative networks are summarized in Table 2.
| Network | Key feature | Use case |
| CNN | Performs convolution operation for feature extraction. | Image enhancement |
| Exhibits outstanding performance in feature extraction. | Image classification and object detection | |
| Captures spatial information of input data efficiently. | Image segmentation | |
| U-Net | Comprises an encoder–decoder structure. | Image enhancement |
| Utilizes skip connections to leverage high-resolution feature maps. | ||
| Demonstrates strong performance even with small data sets. | Image segmentation | |
| Excels in segmentation tasks. | ||
| GAN | Consists of a generator network and a discriminator network. | Image generation |
| Generates data that closely resembles real input data (generator). | ||
| Discriminates between generated data and real data (discriminator). | Image style transfer | |
| Engages in competitive training between the generator and discriminator. | Image/data augmentation | |
| Applies for generating new data. |
Table 2. Three representative networks.
3 Challenges in PAI and Solutions through DL
3.1 Overcoming Limited Detection Capabilities
In PAI, optimal image quality requires a broadband US transducer and dense spatial sampling to enclose the target.43,61,73 However, real-world scenarios introduce limitations, such as limited bandwidth, limited view, and data sparsity. DL methods have been used as postprocessing techniques to overcome these limitations and enhance the PA signals or images, reducing artifacts. This section provides an overview of studies utilizing DL methods as PAI postprocessing methods. The DL-based image reconstruction studies are discussed separately in Sec. 3.4.
3.1.1 Limited bandwidth
The bandwidth of US transducer arrays is limited compared to the natural broadband PA signal (from tens of kilohertz to a hundred megahertz).74 Although optical detectors of PA waves have expanded the detection bandwidth, manufacturing high-density optical detector arrays and adopting them to PACT remains a technical challenge.75
To solve the limited-bandwidth problem, Gutte et al. proposed a DNN with five fully connected layers to enhance the PA bandwidth [Fig. 4(a)].76 The network takes a limited-bandwidth signal as input and outputs an enhanced bandwidth signal, which is then used for PA image reconstruction using DAS. To train the network, the authors generated numerical phantoms using the k-Wave toolbox79 to create pairs of full-bandwidth and limited-bandwidth PA signals. The synthesized results from the numerical phantoms demonstrated an enhanced bandwidth that is like that of images obtained from the full-bandwidth signal [Fig. 4(a)].
![]()
Figure 4.Representative studies using DL methods to overcome limited-detection capabilities. (a) A DNN with five fully connected layers enhances bandwidth. (b) LV-GAN for addressing the limited-view problem. (c) A Y-Net generates the PA images by optimizing both raw data and reconstructed images from the traditional method. (d) A 3D progressive U-Net (3D-pUnet) to diminish the effects of limited-view artifacts and sparsity arising from cluster view detection. The images are adapted with permission from Ref.
3.1.2 Limited view
PA image quality is reduced by the scant information provided by the limited coverage angle of the PA signals detected by the US transducer.3 This problem is commonly encountered in PACT systems, particularly in linear US array-based systems and is referred to as the “limited view” problem.75,80 Researchers have addressed this problem to some extent by developing reconstruction methods with iterative methods.80 Recent studies based on the fluctuation of the PA signal of blood flow or microbubbles81,82 show another effective solution, but they need a number of single images to reconstruct one fluctuation image, which compromises the temporal resolution.
Deng et al.83 developed DL methods using U-Net and principal component analysis processed very deep convolutional networks (PCA-VGG)84 while Zhang et al.85 designed a dual domain U-Net (DuDoUnet) incorporating reconstructed images and frequency domain information. In addition to utilizing the U-Net architecture, researchers have also explored the use of GAN networks, which have garnered attention due to their ability to preserve high-frequency features and prevent oversmoothing in images. Lu et al. proposed a GAN-based network called the limited view GAN (LV-GAN).77Figure 4(b) shows the architecture of LV-GAN, which consists of two networks: the generator network responsible for generating high-quality PA images from limited-view images, and the discriminator network designed to distinguish the generated images from the GT. To ensure accurate and generalizable results, the LV-GAN was trained using both simulated data generated by the k-Wave toolbox and experimental data obtained from a custom-made PA system. The results presented in Fig. 4(b), using ex vivo data, demonstrate the ability of LV-GAN to successfully reconstruct high-quality PA images in limited-view scenarios. The quantitative analysis further confirms that LV-GAN outperforms the U-Net framework, achieving the highest retrieval accuracy.
The combination of a postprocessing method with direct processing using PA signals is considered as another approach to reduce artifacts in limited-view scenarios. Lan et al.78 designed a new network architecture, called Y-net, which reconstructs PA images by optimizing both raw data and reconstructed images from the traditional method [Fig. 4(c)]. This network has two inputs, one from raw PA data and the other from the traditional reconstruction. It combines two encoders, each corresponding to one of the input paths, with a shared decoder path. The training data were generated by the k-Wave toolbox with a linear array setup. The public vascular data set86 was used to generate PA signals. They compared the proposed method with conventional reconstruction methods [e.g., DAS and time reversal (TR)] and other DL methods such as U-Net. In in vitro and in vivo experiments, the proposed method showed superior performance to the other methods, with the best spatial resolution.
3.1.3 Sparsity
To achieve the best image quality, the interval between two adjacent positions of the transducer or array elements must be less than half of the lowest detectable acoustic wavelength, according to the Nyquist sampling criterion.87 In sparse sampling, the actual detector density is lower than this requirement, introducing streak-shaped artifacts in images.74 Sparse sampling can also result from a trade-off between image quality and temporal resolution, which is sometimes driven by system cost and hardware limitations.88
To remove artifacts caused by data sparsity, Guan et al.89 added additional dense connectivity into the contracting and expanding paths of a U-Net. Farnia et al.90 combined a TR method with a U-Net by inserting it in the first layer. Guo et al.91 built a network containing a signal-processing method and an attention-steered network (AS-Net). Lan et al.92 proposed a knowledge infusion GAN (Ki-GAN) architecture that combines DAS and PA signals for reconstruction from sparsely sampled data. DiSpirito et al.93 compared various CNN architectures for PAM image recovery from undersampled data of in vivo mouse brains.94 They chose a fully dense U-Net (FD U-Net) with a dense block, allowing PAM image reconstruction using just 2% of the original pixels. Later, they proposed a new method based on a deep image prior (DIP) method95 to solve this problem without pretraining or GT data.
3.1.4 Combinational limited-detection problems
Previous studies have primarily tackled individual issues in isolation, neglecting the simultaneous occurrence of multiple limited-detection challenges in PA systems.74 However, researchers have recently focused on utilizing a single NN to address two or three limited-detection problems concurrently, leading to promising advancements in this area.
For linear array, Godefroy et al.96 incorporated dropout layers97 into a modified U-Net and further built a Bayesian NN to improve the PA image quality. Vu et al.98 built a Wasserstein GAN (WGAN-GP) that combined a U-Net and a deep convolutional GAN (DCGAN).99 The network reduced limited-view and limited-bandwidth artifacts in PACT images. For a ring-shaped array, Zhang et al.100 developed a 10-layer CNN, termed a ring-array DL network (RADL-net), to eliminate limited-view and under-sampling artifacts in photoacoustic tomography (PAT, also known as PACT) images. Davoudi et al.101 proposed a U-Net network to improve the image quality from sparsely sampled data from a full-ring transducer array. They later updated their U-Net architecture102 to operate on both images and PA signals. Awasthi et al.103 proposed a U-Net architecture to achieve superresolution, denoising, and bandwidth enhancements. They replaced the softmax activation function in the final two layers of the U-Net for segmentation with an exponential linear unit.104 Schwab et al. proposed a network that combined the BP with dynamic aperture length (DAL) correction, which they called DALnet105 to address the limited-view and undersampling issues in the 3D imaging PACT system.
One of the notable achievements in applying DL to the 3D-PACT system was made by Choi et al.52 They introduced a 3D progressive U-Net (3D-pUnet) as a solution to address limited-view artifacts and sparsity caused by clustered-sampling detection, as shown in Fig. 4(d). The design of their network was inspired by the progressive growth GAN,106 which utilizes a progressively increasing procedure to optimize a U-Net. In their 3D-pUnet, subnetworks were trained sequentially using downsampled data from the original high-resolution volume data, gradually transferring knowledge obtained from each progressive step.
The training data set consisted of in vivo experimental data from rats, and the results demonstrated superior performance compared with the conventional 3D-U-Net method. Interestingly, they demonstrated that the 3D-pUnet trained cluster-sampled data set also works in sparsely sampled data sets. The proposed approach was also applied to predict dynamic contrast-enhanced images and functional neuroimaging in rats, achieving increased imaging speed while preserving high image quality. In addition, they demonstrated the ability to accurately measure physiological phenomena and enhance structural information in untrained subjects, including tumor-bearing mice and humans.
All the research reviewed in this section is summarized in Table 3.
| Author | Neural | Basic | Training data set (if specified, | Test data set | Specified task | Representative | |
| Source | Data amount | ||||||
| Gutte et al. | FC-DNN | CNN | Simulation of the breast phantom | 286,300 slices (from 2863 volumes) | Simulation/in vitro phantom | Reduce limited-bandwidth artifacts | CNR (versus DAS) 0.01 → 2.54 |
| PC 0.22 → 0.75 | |||||||
| Deng et al. | U-Net and VGG | U-Net | In vivo mouse liver | 50 | Numerical simulation data/in vitro phantom/in vivo data | Reduce limited-view artifacts from the circular US array | SSIM (versus DAS) 0.39 → 0.91 |
| PSNR 7.54 → 24.34 | |||||||
| Zhang et al. | DuDoUnet | U-Net | k-Wave simulation | 1500 | k-Wave simulation | Reduce limited-view artifacts from the linear US array | SSIM (versus U-Net) 0.909 → 0.935 |
| PSNR 19.4 → 20.8 | |||||||
| Lu et al. | LV-GAN | GAN | k-Wave simulation of absorbers and vessels/in vitro phantom of microsphere and vessel structure | 793 pairs (absorbers)/1600 pairs (vessels)/ | k-Wave simulation of absorbers and vessels/in vitro phantom (microsphere and vessel structure) | Reduce limited-view artifacts from the circular US array | SSIM (versus DAS) 0.135 → 0.871 |
| 30 pairs (microsphere) | PSNR 9.41 → 30.38 | ||||||
| CNR 22.72 → 43.41 | |||||||
| 22 pairs (vessel structures) | |||||||
| Lan et al. | Y-Net | — | k-Wave simulation of segmented blood vessels from DRIVE data set | 4700 | k-Wave simulation/in vitro phantom/in vivo human palm | Reduce limited-view artifacts from the linear US array | SSIM (versus DAS) 0.203 → 0.911 |
| PSNR 17.36 → 25.54 | |||||||
| SNR 1.74 → 9.92 | |||||||
| Guan et al. | FD-UNet | U-Net | k-Wave simulation:/k-Wave simulation of realistic vasculature phantom from micro-CT images of mouse brain | 1000 simulation/1000 (realistic vasculature) | k-Wave simulation:/k-Wave simulation of realistic vasculature phantom (micro-CT images of the mouse brain) | Reduce artifacts from sparse data in the circular US array | SSIM (versus DAS) 0.75 → 0.87 |
| PSNR 32.48 → 44.84 | |||||||
| Farnia et al. | U-Net | U-Net | k-Wave simulation from the DRIVE data set | 3200 | k-Wave simulation from DRIVE data set/in vivo mouse brain | Reduce artifacts from sparse data in the circular US array | SSIM (versus DAS) 0.81 → 0.97 |
| PSNR 29.1 → 35.3 | |||||||
| SNR 11.8 → 14.6 | |||||||
| EPI 0.68 → 0.90 | |||||||
| Guo et al. | AS-Net | Non | k-Wave simulation of human fundus culi vessel/in vivo fish/in vivo mouse | 3600/1744/1046 | k-Wave simulation of human fundus culi vessel/in vivo fish/in vivo mouse | Reduce artifacts from sparse data and speed up reconstruction from the circular US array | SSIM (versus DAS) 0.113 → 0.985 |
| PSNR 8.64 → 19.52 | |||||||
| Lan et al. | Ki-GAN | GAN | k-Wave simulation of retinal vessels from public data set | 4300 | k-Wave simulation of retinal vessels from public data set | Remove artifacts from sparse data from the circular US array | SSIM (versus DAS) 0.215 → 0.928 |
| PSNR 15.61 → 25.51 | |||||||
| SNR 1.63 → 11.52 | |||||||
| DiSpirito et al. | FD U-Net | U-Net | In vivo mouse brain | 304 | In vivo mouse brain | Improve the image quality of undersampled PAM images | SSIM (versus zero fill) 0.510 → 0.961 |
| PSNR 16.94 → 34.04 | |||||||
| MS-SSIM 0.585 → 0.990 | |||||||
| MAE 0.0701 → 0.0084 | |||||||
| MSE 0.0027 → 0.00044 | |||||||
| Vu et al. | DIP | CNN | In vivo blood vessels | — | In vivo blood vessels/non-vascular data | Improve the image quality of undersampled PAM images | SSIM (versus bilinear) 0.851 → 0.928 |
| PSNR 25.6 → 31.0 | |||||||
| Godefroy et al. | U-Net/Bayesian NN | U-Net | Pairs of PAI and photographs of leaves/Corresponded numerical simulation | 500 | PAI and photographs of leaves/numerical simulation | Reduce limited-view and limited-bandwidth artifacts from the linear US array | NCC (versus DAS) 0.31 → 0.89 |
| SSIM 0.29 → 0.87 | |||||||
| Vu et al. | WGAN-GP | GAN | k-Wave simulation: disk phantom and TPM vascular data | 4000 (disk)/7200 (vascular) | k-Wave simulation: disk phantom and TPM vascular data/tube phantom/in vivo mouse skin | Reduce limited-view and limited-bandwidth artifacts from the linear US array | SSIM (versus U-Net) 0.62 → 0.65 |
| PSNR 25.7 → 26.5 | |||||||
| Zhang et al. | RADL-net | CNN | k-Wave simulation | 161,000 (including augmentation and cropping from 126 vascular images) | k-Wave simulation/vascular structure phantom/in vivo mouse brain | Reduce limited-view and sparsity artifacts from the ring-shaped US array | SSIM (versus DAS) 0.11 → 0.93 |
| PSNR 17.5 → 23.3 | |||||||
| Davoudi et al. | U-Net | U-Net | Simulation: planar parabolic absorber and mouse/in vitro circular phantom/in vitro vessel-structure phantom/in vivo mouse | Not mentioned/28/33/420 | Simulation: planar parabolic absorber and mouse/in vitro circular phantom/in vitro vessel-structure phantom/in vivo mouse | Reduce limited-view and sparsity artifacts from the circular US array | SSIM (versus input) 0.281 → 0.845 |
| Davoudi et al. | U-Net | U-Net | In vivo human finger from seven healthy volunteers | 4109 (including validation) | In vivo human finger | Reduce the limited-view and sparsity artifacts from the US circular array | SSIM (versus U-Net) 0.845 → 0.944 |
| PSNR 14.3 → 19.0 | |||||||
| MSE 0.04 → 0.014 | |||||||
| NRMSE 0.818 → 0.355 | |||||||
| Awasthi et al. | Hybrid end-to-end U-Net | U-Net | k-Wave simulation from breast sinogram images | 1000 | k-Wave simulation of the numerical phantom, blood vessel, and breast/ horsehair phantoms/ in vivo rat brain | Super-resolution, denoising, and bandwidth enhancement of the PA signal from the circular US array | PC (versus DAS) 0.307 → 0.730 |
| SSIM 0.272 → 0.703 | |||||||
| RMSE 0.107 → 0.0617 | |||||||
| Schwab et al. | DALnet | CNN | Numerical simulation of 200 projection images from 3D lung blood vessel data | 3000 (after cropping) | Numerical simulation/in vivo human finger | Reduce limited-view, sparsity and limited bandwidth artifacts | SSIM (versus input) 0.305 → 0.726 |
| Correlation 0.382 → 0.933 | |||||||
| Choi et al. | 3D-pUnet | U-Net | In vivo rat | 1089 | In vivo rat/in vivo mouse/in vivo human | Reduce limited-view and sparsity artifacts | MS-SSIM (versus input) 0.83 → 0.94 |
| PSNR 32.0 → 34.8 | |||||||
| RMSE 0.025 → 0.019 |
Table 3. Summary of overcoming the limited detection capabilities with DL approaches.
3.2 Compensating for Low-Dosage Light Delivery
Pulsed laser sources, such as an optical parametric oscillator laser system with a Nd:YAG pumped laser, are commonly used in PACT systems to achieve deep penetration with a high SNR, but those laser systems are bulky and expensive.2,107 In recent years, researchers have explored compact and less expensive alternatives, such as pulsed-laser diodes107 and light-emitting diodes (LEDs).108 While these alternatives have shown promising results, their low pulse energy results in a low SNR, requiring frame averaging to increase image quality. Unfortunately, this method comes at a cost, as it reduces imaging speed. Furthermore, in dynamic imaging, frame averaging can cause blurring or ghosting due to the movement of the object being imaged. To address these problems, DL methods can be applied to enhance image quality in situations where the light intensity is low.
One of the representative works for LED-based systems was achieved by Hariri et al.53 They proposed a multilevel wavelet-convolutional NN (MWCNN) that could map the low-fluence PA images to high-fluence PA images from an Nd:YAG laser system. This approach helps to eliminate the background noise while preserving the structures of the target, as shown in Fig. 5(a). Phantom and in vivo studies were conducted to assess the performance of their model. The MWCNN demonstrated a significant improvement in contrast-to-noise ratio (CNR) with up to a 4.3-fold enhancement in the phantom study and a 1.76-fold enhancement in the in vivo study. These results highlight the practicality of the proposed method in real-world scenarios. Singh et al.111 and Anas et al.112 proposed a U-Net and a deep CNN-based approach to improve the image quality with a similar system setup. Anas et al. later introduced a recurrent neural network (RNN)113 to further improve the system’s performance.114
![]()
Figure 5.Representative DL approaches compensate for low laser dosage. (a) An MWCNN that generates high-quality PA images from low-fluence PA images. (b) An HD-UNet that enhances the image quality in a pulsed-laser diode PACT system. (c) An MT-RDN that performs image denoising, superresolution, and vascular enhancement. The images are adapted with permission from Ref.
To enhance the image quality in a pulsed-laser-diode PA system, Rajendran et al.109 proposed a hybrid dense U-Net (HD-UNet) [Fig. 5(b)]. To train the network, they generated simulated data using the k-Wave toolbox, and evaluated the model with both single- and multi-US transducer (1-UST and multi-UST-PLD) PACT systems, using both phantom and in vivo images. Compared with their previous system, the HD-UNet improved the imaging speed by approximately 6 times in the 1-UST system and 2 times in the multi-UST-PLD system. To address the challenges of balancing laser dosage, imaging speed, and image quality in OR-PAM, Zhao et al.110 proposed a multitask residual dense network (MT-RDN) that performs image denoising, superresolution, and vascular enhancement [Fig. 5(c)]. The network comprises three subnetworks, each using an independent RDN framework and assigned a supervised learning task. The first subnetwork processes the data of input 1 (i.e., 532 nm data) to obtain output 1, and the second subnetwork processes the data of input 2 (i.e., 560 nm data) to obtain output 2. These outputs are then combined and processed by subnetwork 3, and the differences between the outputs and the GT are compared.
To train the network, input images were undersampled at half-per-pulse laser energy of the GT, while the GT images were sampled at the full ANSI per-pulse fluence limit. To evaluate the performance of the proposed method, U-Net and RDN were used. The MT-RDN method achieved a 16-fold reduction in laser dosage at 2 times data undersampling and a 32-fold reduction in dosage at 4 times undersampling compared to the GT images.
All the research reviewed in this section is summarized in Table 4.
| Author | Neural | Basic | Training data set (if specified, | Test data set | Specified task | Representative evaluation results | |
| Source | Data | ||||||
| Hariri et al. | MWCNN | U-Net | Agarose hydrogel phantom: LED-based PA image and Nd:YAG-based PA image | 229 | Agarose hydrogel phantom of LED-based PA image/in vivo mouse | Denoise PA images from low-dosage system | SSIM (versus input) 0.63 → 0.93 |
| PSNR 15.58 → 53.88 | |||||||
| Singh et al. | U-Net | U-Net | LED-based and Nd:YAG-based tube phantom | 150 | LED-based phantom using ICG and MB | Reduce the frame averaging | SNR 14 → 20 |
| Anas et al. | — | CNN | In vitro phantom | 4536 | In vivo fingers | Improve the quality of PA images | SSIM (versus average) 0.654 → 0.885 |
| PSNR 28.3 → 36.0 | |||||||
| Anas et al. | — | CNN and LSTM | In vitro wire phantom/in vitro nanoparticle phantom | 352,000 | In vitro phantom/in vivo human fingers | Improve the quality of PA images | SSIM (versus input) 0.86→0.96 |
| PSNR 32.3 → 37.8 | |||||||
| Rajendran et al. | HD-Unet | U-Net | k-Wave simulation | 450 | In vitro phantom/in vivo rat | Improve the frame rate | SSIM (versus U-Net) 0.92 → 0.98 |
| PSNR 28.6 → 32.9 | |||||||
| MAE 0.025 → 0.017 | |||||||
| Zhao et al. | MT-RDN | — | In vivo mouse brain and ear | 6696 | In vivo mouse brain and ear | Improve the quality from low dosage laser and downsampled data | SSIM (versus input) 0.64 → 0.79 |
| PSNR 21.9 → 25.6 |
Table 4. Summary of studies on compensating for low-dosage light delivery.
3.3 Improving the Accuracy of Quantitative PAI
Quantitative photoacoustic imaging (qPAI) quantifies molecular concentrations in biological tissue using multiwavelength PA images, enabling the estimation of various endogenous and exogenous contrast agents and physiological parameters, such as
Researchers have pursued multiple avenues to extract fluence distribution information from multiwavelength PA images using DL architectures. Cai et al.119 introduced ResU-Net, which adds a residual learning mechanism to the U-Net. Chang et al.120 developed DR2U-Net, a fine-tuned deep residual recurrent U-Net. Luke et al.121 combined two U-Nets to create a new network called O-Net, which segments blood vessels and estimates
One of the representative results is from Ref. 124. Researchers built two separated convolutional encoder–decoder type networks with skip connections, termed EDS to solve this problem in 3D conditions [Fig. 6(a)]. One network was trained to output images of
![]()
Figure 6.Representative studies to improve the accuracy of quantitative PAI by DL. (a) Convolutional encoder–decoder type network with skip connections (EDS) to produce accurate estimates of
Researchers have also employed DL methods to recover the absorption coefficient from reconstructed PA images. Chen et al.126 proposed a U-Net-based DL network to recover the optical absorption coefficient and Grohl et al.127 adapted a U-Net to compute error estimates for optical parameter estimations. A notable contribution was made by Li et al. in a recent study.54 They addressed the challenge of insufficient data-label pairs in qPAI by introducing two DNNs, depicted in Fig. 6(b). First, they introduced a simulation-to-experiment end-to-end data translation network (SEED-Net) that provides GT images for experimental images through unsupervised data translation from a simulation data set. They then designed a dual-path network based on U-Net (QPAT-Net) to reconstruct images of the absorption coefficient for deep tissues. The QPAT-Net outperformed the previous QPAT method128 in simulation, ex vivo, and in vivo, with more accurate absorption information and relatively few errors.
Another seminal study was done by Zou et al.125 They developed the US-enhanced U-Net model (US-Unet), which combines information from US images and PA images to reconstruct the optical absorption distribution [Fig. 6(c)]. They implemented a pretrained ResNet-18 to extract features from US images of ovarian lesions.
This feature information was incorporated into a U-Net structure designed to reconstruct the optical absorption coefficient. The U-Net was trained on simulation data and subsequently tested on a phantom, blood tubes, and clinical data from 35 patients. The US-Unet outperformed both the U-Net model without US features and the standard DAS method in phantom and clinical studies, demonstrating its potential for improving accuracy in clinical PAI applications.
Compensating for the distribution of light fluence can improve the accuracy of qPAI.129
All the research reviewed in this section is summarized in Table 5.
| Author | Neural | Basic | Training data set | Test data set | Specified task | Representative | |
| Source | Data amount | ||||||
| Cai et al. | ResU-net | U-Net | Numerical simulation | 2048 | Numerical simulation | Extract information from multispectral PA images | Relative errors (versus linear unmixing) 36.9% → 0.76% |
| Chang. et al. | DR2U-net | U-Net | Monte Carlo simulation of simulated tissue structure | 2560 | Monte Carlo simulation of simulated tissue structure | Extract fluence distribution from optical absorption images | Relative Errors (versus linear unmixing) 48.76% → 1.27% |
| Luke et al. | O-Net: | U-Net | Monte Carlo simulation of epidermis, dermis, and breast tissue | 1600 pairs (one pair has two-wavelength PA data) | Monte Carlo simulation of epidermis, dermis, and breast tissue | Estimate the oxygen saturation and segment | Relative errors (versus linear unmixing) 43.7% → 5.15% |
| Yang et al. | EDA-net | — | Monte Carlo and k-Wave simulation from female breast phantom | 4888 | Monte Carlo and k-Wave simulation based on clinically obtained female breast phantom | Extract the information from the multi-wavelength PA images | Relative errors (versus linear unmixing) 41.32% → 4.78% |
| Gröhl et al. | Nine-layer fully connected NN | CNN | Monte Carlo simulation of in silico vessel phantoms | 776 | In vivo porcine brain and human forearm | Obtain quantitative estimates for blood oxygenation | No statistical results |
| Bench et al. | EDS | U-Net | k-Wave simulation of human lung from lung CT scans/k-Wave simulation of three-layer skin model | — | k-Wave simulation | Produce 3D maps of vascular | Mean difference (versus linear unmixing) 6.6% → 0.3% |
| Chen et al. | U-Net | U-Net | Monte Carlo simulation | 2880 | In vitro phantom | Recover the optical absorption coefficient | Relative error less than 10% |
| Gröhl et.al | U-Net | U-Net | Monte Carlo and k-Wave simulations of in silico tissue | 3600 | Monte Carlo and k-Wave simulation of in silico | Improve optical absorption coefficient estimation | Estimation error (versus linear unmixing) 58.3% → 3.1% |
| Li et al. | Two GANs: SEED-Net and QOAT-Net | GAN | Numerical simulation of phantom, mouse, and human brain/experimental data of phantom, ex vivo, and in vivo mouse | 3040, 2560, and 2560/2916, 3200, and 3800 | Ex vivo porcine tissue, mouse liver, and kidney/In vivo mouse | Improve optical absorption coefficient estimation | Relative errors (versus linear unmixing) 8.00% → 4.82% |
| Relative errors 8.00% → 4.82% | |||||||
| Zou et al. | US-Unet | U-Net | Monte Carlo and k-Wave simulation/in vitro phantom | 2000/480 | In vitro blood tube/in vivo clinical data set | Improve optical absorption coefficient estimation | Accuracy (versus linear unmixing) 0.71 → 0.89 |
| Madasamy et al. | Network comparing: U-Net, FD U-Net, Y-Net, FD Y-Net, Deep ResU-Net, and GAN | — | 2D numerical simulation of retinal fundus (from Kaggle and RFMID)/3D numerical simulation of breast phantom | 1858 (before augmentation)/5 3D volumes (12,288 slices after augmentation) | 2D numerical blood vessel/3D numerical breast phantom | Fluence correction | PSNR (versus linear unmixing) 37.9 → 45.8 |
| SSIM 0.80 → 0.96 | |||||||
| Durairaj et al. | Two networks: initialization network and unmixing network | — | NIRFAST and k-Wave simulation | Not mentioned | NIRFAST and k-Wave simulation | Unmix the spectral information | Regardless of prior spectral information |
| Olefir et al. | DL-eMSOT: bi-directional RNN with two LSTMs | — | Monte Carlo simulation | 10,944 | In vitro phantom/in vivo mouse | Replace inverse problem of eMSOT | Mean error (versus eMSOT) 4.9% → 1.4% |
| Median error 3.5% → 0.9% | |||||||
| Standard deviation 4.8% → 1.5% |
Table 5. Summary of studies to improve the accuracy of quantitative PAI.
3.4 Optimizing or Replacing Conventional Reconstruction Algorithms
In PACT, the acoustic inverse problem involves reconstructing the PA initial pressure from raw data. Several reconstruction methods have been developed, including BP,47 FB,48 DAS,44 DMAS,46 TR,49 and model-based methods.51 However, each method has limitations, and either to enhance existing reconstruction techniques or to directly reconstruct PA images using NNs, researchers have turned to DL methods.
Various DL methods have been developed to convert PA raw data into images. One such method, called Pixel-DL, proposed by Guan et al.,137 uses pixel-wise interpolation followed by an FD U-Net for limited-view and sparse PAT image reconstruction [Fig. 7(a)]. The Pixel-DL model was trained and tested using simulated PA data from synthetic, mouse brain, lung, and fundus vasculature phantoms. It achieved comparable or better performance than iterative methods and consistently outperformed other CNN-based approaches for correcting artifacts.
![]()
Figure 7.Representative studies to optimize conventional reconstruction algorithms or replace them with DL. (a) Pixel-wise interpolation approach followed by an FD-UNet for limited-view and sparse PAT image reconstruction. (b) End-to-end U-Net with residual blocks to reconstruct PA images. (c) Two-step PA image reconstruction process with FPnet and U-Net. The images are adapted with permission from Ref.
To direct reconstruct PA images, Waibel et al.139 introduced a modified U-Net that includes additional convolutional layers in each skip connection. Antholzer et al.140 proposed a direct reconstruction process, based on a U-Net and a simple CNN, that can resolve limited-view and sparse-sampling issues. Lan et al.141 proposed a modified U-Net, termed DU-Net, to reconstruct PA images using multifrequency US-sensor raw data. A noteworthy study of this topic is an end-to-end reconstruction network developed by Feng et al.,138 termed Res-U-Net [Fig. 7(b)]. They integrated residual blocks into the contracting and symmetrically expanding path of U-Net and added a skip connection between the input of raw data and the output of images. The training, validation, and test data sets were synthesized using the k-Wave toolbox. In digital phantom experiments, the Res-UNet showed performance superior to other reconstruction methods [Fig. 7(b)].
Another representative work was done by Tong et al.55 They proposed a novel two-step reconstruction process with a feature projection network (FPnet) and a U-Net [Fig. 7(c)]. The FPnet converts PA signals to images and contains several convolutional layers to extract features. There is one max pooling layer for downsampling and one full connection layer for domain transformation. The U-Net performs postprocessing to improve image quality. The resulting network, trained using numerical simulations and in vivo experimental data, outperformed other approaches to handle limited-view and sparsely sampled experimental data, exhibiting superior performance on in vivo experiments [Fig. 7(c)].
In addition, Yang et al.142 introduced recurrent inference machines (RIM), an iterative PAT reconstruction method using convolution layers. Kim et al.143 employed upgUNET, a U-Net model with 3D transformed arrays for image reconstruction. Hauptmann et al.144 proposed DGD, a deep gradient descent algorithm, outperforming U-Net and other model-based methods. They also introduced fast-forward PAT (FF-PAT), a modified version of DGD, which addressed artifacts using a small multiscale network.145
All the research reviewed in this section is summarized in Table 6.
| Author | Neural | Basic | Training data set (if specified, validation is excluded) | Test data set | Specified task | Representative | |
| Source | Data amount | ||||||
| Guan | Pixel-DL | U-Net | k-Wave simulation: circles, Shepp-Logan, and vasculature, vasculature phantom from micro-CT images of mouse brain | 1000 (circles),/1000 (realistic vasculature) | k-Wave simulation: circles, Shepp-Logan, and vasculature, vasculature phantom from micro-CT images of mouse brain | Reconstruct PA images | PSNR (versus TR) |
| SSIM 0.52 → 079 | |||||||
| Waibel | U-Net | U-Net | Monte Carlo and k-Wave simulation | 2304 | Monte Carlo and k-Wave simulation | Reconstruct PA images | IQR (versus DAS) |
| Antholzer | U-Net | U-Net | Numerical simulation of ring-shaped phantoms | 1000 | Numerical simulation of ring-shaped phantoms | Reconstruct PA images | MSE (versus general |
| Lan | DU-Net | U-Net | k-Wave simulation: disc phantom and segmented fundus oculi/vessels CT | 4000 | k-Wave simulation: disc phantom and segmented fundus oculi/ vessels CT | Reconstruct PA images | PSNR (versus DAS) |
| SSIM 0.394 → 0.994 | |||||||
| Feng | Res-UNet | U-Net | k-Wave simulation: disc bread, spider (from “quick draw”), simple wires, logos, natural phantom | 58,126 (80% of | k-Wave simulation: disc, PAT, vessel/in vitro phantom | Reconstruct PA images | Vessel phantom |
| PC (versus MRR) | |||||||
| PSNR 6.57 → 13.29 | |||||||
| Tong | FPnet+ | U-Net | Numerical simulation: brain from MRI, abdomen from MRI, vessel from DRIVE data set/in vivo mouse brain and abdomen | 15,757: 2211 (brain), | Numerical simulation: brain, abdomen and liver cancer from MRI, vessel/ | Reconstruct PA images | MSOT-Abdomen |
| PSNR (versus FBP) | |||||||
| SSIM 0.2647 → 0.9073 | |||||||
| RMSE 0.4771 → 0.0910 | |||||||
| Yang | RIM | — | k-Wave simulation of segmented blood vessels from DRIVE data set | 2400 | k-Wave simulation of segmented blood vessels from DRIVE | Reconstruct PA images | PSNR (versus DGD) |
| Kim | upgUNET | U-Net | Monte Carlo simulation | 128,000 (after | Monte Carlo simulation/ | Reconstruct PA images | PSNR (versus DAS) |
| SSIM 0.208 → 0.754 | |||||||
| Hauptmann | Updated | DGD | k-Wave simulation pf human lung from 50 | 1024 (from 50 | k-Wave simulation pf human lung from 50 | Reconstruct PA images | PSNR (versus U-Net) |
| SSIM 0.933 → 0.945 | |||||||
| Hauptmann | FF-PAT | U-Net | k-Wave simulation of human lung from lung | 1024 (from 50 | k-Wave simulation of human lung/in vivo data | Reconstruct PA images | PSNR (versus BP) |
Table 6. Summary of methods to optimize or replace conventional reconstruction algorithms.
3.5 Addressing Tissue Heterogeneity
Biological tissues are acoustically nonuniform, making it crucial to use a locally appropriate speed of sound (SoS) value for accurate PA reconstruction. SoS mismatch or a discontinuity in hard textured tissue can create acoustic reflection and imaging artifacts74 that make it hard to detect the source of the PA signal, which is especially troublesome for interventional applications. DL methods have been used to detect point sources, remove reflections, and mitigate the difficulties presented by acoustic heterogeneity.
Highly echogenic structures can cause a reflection of a PA wave to appear to be a true signal,146 which makes it hard to find point targets or real sources in PAI. Reiter et al.147 trained a CNN to identify and remove reflection noise, locate point targets, and calculate absorber sizes in PAI. Later, Allman et al.148 found Fast-RCNN149 to be more effective than VGG1684 for source detection and artifact elimination. Shan et al.150 incorporated a DNN into an iterative algorithm to correct reflection artifacts, achieving superior results compared with other methods.151,152
Jeon et al.56 proposed a generalized solution to mitigate SoS aberration in heterogeneous tissue by DL. They proposed a hybrid DNN model, named SegU-net, based on U-Net and SegNet153 [Fig. 8(a)]. The architecture is similar to SegNet, but has an additional connection between the encoder and decoder through concatenation layers, like U-Net. The training data were generated using the k-Wave toolbox with different SoS values. They tested the model with phantoms with homogeneous media and in heterogeneous media. The proposed method showed better results than the multistencil fast marching155 method and automatic SoS selection.156 It not only resolved the SoS aberration but also removed streak artifacts in images of healthy human limbs and melanoma.
![]()
Figure 8.Representative DL studies to correct the SoS and improve the accuracy of image classification and segmentation. (a) Hybrid DNN model including U-Net and Segnet to mitigate SOS aberration in heterogeneous tissue. (b) Sparse-UNet (S-UNet) for automatic vascular segmentation in MSOT images. The images are adapted with permission from Ref.
All the research reviewed in this section is summarized in Table 7.
| Author | Neural network | Basic | Training data set | Test data set | Specified task | Representative | |
| Source | Data | ||||||
| Reiter et al. | CNN | CNN | k-Wave simulation | 19,296 | k-Wave simulation/in vitro vessel-mimicking target phantom | Identify point source | — |
| Allman et al. | CNN consisting of VGG16/Fast R-CNN | CNN | k-Wave simulation | 15,993 | k-Wave simulation/in vivo data | Identify and remove reflection artifacts | Precision, recall, and AUC > 0.96 |
| Allman et al. | CNN consisting of VGG16/fast R-CNN | CNN | k-Wave simulation | 15,993 | In vitro phantom | Correct reflection artifact | Accuracy (phantom) 74.36% |
| Shan et al. | U-Net | U-Net | Numerical simulation from 3 cadaver CT | 64,000 | Numerical simulation from 1 cadaver CT | Correct reflection artifacts | PSNR (versus TR) 9 → 29 |
| SSIM (versus TR) 0.2 → 0.9 | |||||||
| Jeon et al. | SegU-net | U-Net | k-Wave simulation of in silico phantom | 270 | k-Wave simulation of in silico phantom/in vivo human forearm and foot | Reduce speed-of-sound aberrations | In silico phantom |
| SSIM (versus pre-corrected) + 0.24 |
Table 7. Summary of methods for addressing tissue heterogeneity.
3.6 Improving the Accuracy of Image Classification and Segmentation
As PAI gains increasing attention in clinical studies, more accurate classification and segmentation methods are necessary to improve the interpretation of PA images. Image segmentation extracts the outline of objects within an image and identifies the distinct parts of the image that correspond to these objects.158,159 Image classification predicts a label for an image, identifying its content. In this section, we focus on segmentation and classification techniques.158
Segmentation and classification are widely used in image postprocessing, and transfer learning has been utilized to take advantage of pretrained DNNs. Zhang et al.160 used DL models AlexNet161 and GoogLeNet162 for PA image classification, outperforming support vector machine (SVM). Jnawali et al.163 employed transfer learning with Inception-ResNet-V2164 for thyroid cancer detection and introduced a deep 3D CNN for cancer detection in multispectral photoacoustic data sets.165 Moustakidis et al.166 developed SkinSeg for identifying skin layers in raster-scan optoacoustic mesoscopy (RSOM) images, evaluating decision trees,167 SVM,168 and DL algorithms. Nitkunanantharajah et al.169 achieved good classification performance with ResNet18164 on RSOM nail fold images.
PACT has demonstrated its great potential for human vascular imaging in several clinical studies.15 However, segmentation of the vascular structures, particularly the vascular lumen, is still accomplished through manual delineation by expert physicians, which is not only time-consuming but also subjective. To address this issue, Chlis et al.154 proposed a sparse UNet (S-UNet) for automatic vascular segmentation on MSOT images. GT is obtained from binary images extracted from MSOT images, based on consensus between two clinical experts [Fig. 8(b)]. The MSOT raw data from six healthy humans’ vasculature were acquired using a handheld MSOT system, and they were split into training, validation, and test sets. The S-Unet showed performance similar to other U-Net methods, but with smaller parameter sizes and the ability to select wavelengths, indicating its potential for clinical application.170
In addition, Lafci et al.171 proposed a U-Net architecture to accurately segment animal boundaries in hybrid PA and US (PAUS) images. Boink et al.172 proposed a learned primal-dual (L-PD) algorithm based on a CNN to solve the reconstruction and segmentation problem simultaneously. Ly et al.57 introduced a modified U-Net DL model for automatic skin and vessel segmentation in in vivo PAI. The U-Net architecture showed the best performance.
All the research reviewed in this section is summarized in Table 8.
| Author | Neural | Basic | Training data set (if specified, | Test data set | Specified task | Representative | |
| Source | Data amount | ||||||
| Zhang et al. | AlexNet/ | CNN | k-Wave simulation from | 98 (normal)/ | k-Wave simulation from | Classify and segment | BI-RADS rating accuracy: |
| Jnawali et al. | Inception- | — | Ex vivo human thyroid | 73 | Ex vivo human thyroid | Detect cancer tissue | AUCs for cancer, benign, |
| Jnawali et al. | 3D CNN | CNN | Thyroid cancer tissue | 74 (thyroid)/ | Thyroid and prostate | Detect cancer tissue | AUC, 0.72 |
| Moustakidis | SkinSeg | — | In vivo human | About 26,190 | In vivo human | Identify skin | Per-class accuracy: |
| Nitkunanantharajah | ResNet18 | — | In vivo human nailfold | 990 (from 33 subjects) | In vivo human nailfold | Classify images | Intra-class correlation: |
| Sensitivity and specificity: | |||||||
| Chlis et al. | S-Unet | U-Net | In vivo human | 98 pairs (one pair has | In vivo human | Segment human | Dice coeff. (versus U-Net): |
| Schellenberg | nnU-Net | U-Net/ | PA/US images of | 144 PA and US | 36 images for validation | Segment images | Dice coeff. (versus FCNN): |
| Normalized surface | |||||||
| Lafci et al. | U-Net | U-Net | In vivo mice brain, | 174 images from | In vivo mice brain, | Segment hybrid | Dice coeff. 0.95 |
| Boink et al. | L-PD | — | Retinal blood vessels | 768 | Retinal blood vessels | Reconstruct and | PSNR (versus FBP): |
| Ly et al. | Modified | U-Net | In vivo human palm | 800 | In vivo human palm | Segment blood | Global accuracy: |
| Sensitivity: 0.6406 (SegNet-5), |
Table 8. Summary of methods for improving the accuracy of image classification and segmentation.
3.7 Overcoming Other Specified Issues
In addition to the general challenges to PAI mentioned above, researchers encounter several more specific problems with their imaging systems. Collectively, these specific issues constitute a seventh challenge, and among them we have identified six representative categories: motion artifacts, limited spatial resolution, electrical noise and interference, image misalignment, slow accelerating superresolution imaging, and achieving digital histologic staining.
Motion artifacts caused by breathing or heartbeats can significantly reduce image quality in PAM and PA endoscopy (PAE or intravascular PA, IVPA). To address this issue, researchers have presented various breathing artifact removal methods,173,174 and DL methods have recently been proposed as a potential solution. Chen et al.175 introduced a CNN approach with three convolutional layers to address motion artifacts and pixel dislocation in in vivo rat brain images. Zheng et al.176 proposed MAC-Net, a network based on VGG16 GAN134 and spatial transformer networks (STN),177 to suppress motion artifacts in IVPA. Both methods demonstrated successful improvement in image quality.
OR-PAM can penetrate
![]()
Figure 9.Representative studies using DL to solve specific issues. (a) GAN-based framework (Wasserstein GAN) to enhance the spatial resolution of AR-PAM. (b) GAN with U-Net to reconstruct superresolution images from raw image frames. (c) Deep-PAM generates virtually stained histological images for both thin sections and thick fresh tissue specimens. The images are adapted with permission from Ref.
DL methods have been applied to address noise and interference issues in PA imaging. Dehner et al.182 developed a discriminative DNN using a U-Net architecture to separate electrical noise from PA signals, improving PA image contrast and spectral unmixing performance. He et al.183 proposed an attention-enhanced GAN with a modified U-Net generator to remove noise from PAM images, prioritizing fine-feature restoration. Gulenko et al.184 evaluated different CNN architectures and found that U-Net demonstrated higher efficiency and accuracy in removing electromagnetic interference noise from PAE systems.
To address image misalignment in PAM, Kim et al.185 utilized a U-Net framework. Their method effectively addressed nonlinear mismatched cross-sectional B-scan PA images during bidirectional raster scanning, resulting in a significant improvement in imaging speed, doubling the speed compared to conventional approaches.
To improve the temporal resolution of superresolution localization imaging,186,187 hundreds of thousands of overlapping images are traditionally required. However, this process can be time-consuming. To address this problem, Kim et al.180 proposed a GAN with U-Net based on pix2pix188 to reconstruct superresolution images from raw image frames [Fig. 9(b)]. The proposed network can be applied to both 3D label-free localization OR-PAM and 2D labeled localization PACT. The authors trained and validated the network with in vivo data from 3D OR-PAM and 2D PACT images. The proposed method reduced the required number of raw frames by 10-fold for OR-PAM and 12-fold for PACT, resulting in a significant improvement in temporal resolution.
Ultraviolet PAM (UV-PAM) takes advantage of the optical absorption contrast of UV light to highlight cell nuclei, generating PA contrast images similar to hematoxylin and eosin (H&E) labeling.189 DL techniques can be used to digitally generate histological stains using trained NNs based on UV-PAM images, providing label-free alternatives to standard chemical staining methods.190
Boktor et al.191 utilized a DL approach based on GANs to digitally stain total-absorption PA remote sensing (TA-PARS) images, achieving high agreement with the gold standard of histological staining. Cao et al.192 employed a cycle-consistent adversarial network (CycleGAN)193 model to virtually stain UV-PAM images, producing pseudo-color PAM images that matched the details of corresponding H&E histology images.
In a recent study, Kang et al.58 combined UV-PAM with DL to generate rapid and label-free histological images [Fig. 9(c)]. Their proposed method, termed deep-PAM, can generate virtually stained histological images for both thin sections and thick fresh tissue specimens. By utilizing an unpaired image-to-image translation network, a CycleGAN, they were able to process GM-UV-PAM images and instantly produce H&E-equivalent images of unprocessed tissues. This groundbreaking approach has significant implications for the field of histology and may offer an alternative to traditional staining methods.
All the research reviewed in this section is summarized in Table 9.
| Author | Neural | Basic | Training data set (if specified; | Test data set | Specified task | Representative evaluation | |
| Source | Amount | ||||||
| Chen | CNN | CNN | — | — | Simulation/in vivo rat brain | Correct motion artifact | — |
| Zheng | MAC-Net | GAN | Simulation | 7680 | Simulation/in vivo IVUS and IVOCT | Correct motion artifact | AIFD (versus pre-corrected) |
| Cheng | WGAN-GP | GAN | In vivo mouse ear | 528 | In vivo mouse ear | Transform AR-PAM | PSNR (versus blind deconv.) |
| SSIM 0.27 → 0.61 | |||||||
| PC 0.76 → 0.78 | |||||||
| Zhang | AOTDL-GAN/ | GAN/U-Net | Simulation/in vitro | 3500 | Simulation/in vitro phantom/in vivo mouse | Transform AR-PAM | SNR (versus Deconv) |
| CNR 6.93 → 12.50 | |||||||
| Lateral resolution | |||||||
| Dehner | U-Net | U-Net | Simulated PA image/Pure | 3000/2110/— | Simulated PA image/Pure electrical noise/phantom/in vivo human breast/simulated white Gaussian noise | Remove noise | SNR of sinograms |
| He | GAN | GAN | Leaf phantom/in vivo | 236/149 | Leaf phantom/in vivo mouse ear | Remove noise | SNR (versus input) |
| CNR 4.80 → 7.63 | |||||||
| Kim | MS-FD-U-Net | GAN | In vivo mouse ear | 830 | In vivo mouse ear | Align bidirectional | SSIM (versus input) |
| PSNR 50.22 → 50.62 | |||||||
| MSE 1.09 → 0.99 | |||||||
| Gulenko | Modified U-Net | U-Net | In vivo rat colorectum/ | 700 | In vivo rat colorectum/in vivo rabbit transurethral | Remove noise | Log(RMSE) (versus Segnet) |
| Log(SSIM) −2.2 → -0.25 | |||||||
| Log(MAE) 2.6 → 2.0 | |||||||
| Kim et al. | U-Net | U-Net | In vivo OR-PAM mouse | 3000/500 | In vivo OR-PAM mouse ear/in vivo PACT mouse brain | Accelerate localization | PSNR (versus input) |
| MS-SSIM 0.89 → 0.97 | |||||||
| Boktor | Pix2Pix GAN | GAN | Experiments | 15,000 | Experiments | Perform virtual staining | SSIM between H&E |
| Cao | CycleGAN | GAN | Experiments | 17,940 (UV-PAM)/26,565 (H&E) | Experiments | Perform virtual staining | H&E versus UV-PAM |
| Cell count: 5549 and 5423. | |||||||
| Nuclear area ( | |||||||
| Internuclear dist.: | |||||||
| Kang | CycleGAN | GAN | Experiments | 400 (thin section)/800 (thick and fresh tissue) | Experiments | Perform virtual staining | H&E versus UV-PAM |
| Cell count: 289 and 283. | |||||||
| Nuclear area ( |
Table 9. Summary of methods for addressing other specified issues.
4 Discussion and Conclusion
PAI is a rapidly growing biomedical imaging modality that utilizes endogenous chromophores to noninvasively provide biometric information, such as vascular structure and
CNN, U-Net, and GAN have been the most representative networks used in PAI-related research. While some studies use basic architectures to achieve their goals, others modify or develop new architectures to solve particular problems in PAI. These networks can be used in various ways, such as postprocessing reconstructed images with different types of noise or directly reconstructing PA images from the time domain in the image domain.
Furthermore, recent research has aimed to extract more accurate quantitative information by using multiple networks, rather than solely focusing on enhancing image quality with one network. This approach can provide more comprehensive and detailed information, improving the overall performance of PAI. While SSIM is commonly used as the loss function, other metrics, such as PSNR and the Pearson correlation, may be added to improve information extraction and convergence speed. The continued exploration and refinement of these network architectures and loss functions will likely contribute to continued advancements in PAI.
Several obstacles remain. The success of DL approaches in PAI is highly dependent on the availability of high-quality data sets, and there is a scarcity of experimental training data. DL approaches in PAI also lack a standardized PA-image format and publicly available data that are accessible to all groups. Consequently, researchers rely on data generated from experiments or simulations, and even publishing PA data is difficult because there is no standard format. The k-Wave79 toolbox is the most generally used to generate the PA initial pressure, along with other light transport simulators, such as mcxlab,194 to generate the light distribution. However, creating reliable simulation data requires GT data from the real world. Commonly, x-ray CT or MRI images of blood vessels and organs are used for PA simulation. Fortunately, there are public data sets of x-ray CT and MRI images, and many groups have used these open data sets to generate PA GTs. However, the varying information obtained by different imaging modalities may not align with PAI. Recently, the International Photoacoustic Standardization Consortium (IPASC)195 has been working to overcome this challenge by bringing together researchers, device developers, and government regulators to achieve standardization of PAI through community-led consensus building. With the efforts and involvement of IPASC, the generalization ability of DL, which is the fundamental problem in medical imaging field, will increase.
While DL has shown improved image qualities in PAI, there are still concerns regarding its applications in biomedical images. Therefore, the efforts of researchers who aim to advance PAI without applying DL are still valuable. For examples, new restoration algorithms196 are being developed to enhance the image quality affected by limited-detection capabilities. The development of ultrawide detection bandwidth transducers197 aims to mitigate the limited bandwidth of traditional US transducers, thereby improving the overall sensitivity and resolution of PAI. Furthermore, specially designed PACT systems with fast-sweep laser scanning techniques offer automatic fluence compensation and motion correction.129 Combining PACT with transmission-mode US-computed tomography enables the mapping of the distribution of SoS, further enhancing PACT image quality.198 Moreover, a wide range of exogenous contrast agents has been developed to improve the SNR of PAI or to overcome the resolution limitations.4
Despite the challenges faced in applying DL to PAI, there is no doubt that DL will have a great impact on the biomedical imaging field, well beyond PAI.199
Biographies of the authors are not available.
References
[31] J. Yao, L. V. Wang. Photoacoustic microscopy. Laser Photonics Rev., 7, 758-778(2013).
[64] S. Ioffe, C. Szegedy. Batch normalization: accelerating deep network training by reducing internal covariate shift, 448-456(2015).
[67] S. Ruder. An overview of gradient descent optimization algorithms(2016).
[70] J. Long, E. Shelhamer, T. Darrell. Fully convolutional networks for semantic segmentation, 3431-3440(2015).
[71] I. Goodfellow et al. Generative adversarial networks. Commun. ACM, 63, 139-144(2020).
[72] D. Berthelot, T. Schumm, L. Metz. BEGAN: boundary equilibrium generative adversarial networks(2017).
[84] K. Simonyan, A. Zisserman. Very deep convolutional networks for large-scale image recognition(2014).
[85] J. Zhang et al. Limited-view photoacoustic imaging reconstruction with dual domain inputs based on mutual information, 1522-1526(2021).
[97] Y. Gal, Z. Ghahramani. Dropout as a Bayesian approximation: representing model uncertainty in deep learning, 1050-1059(2016).
[104] D.-A. Clevert, T. Unterthiner, S. Hochreiter. Fast and accurate deep network learning by exponential linear units (ELUS)(2015).
[105] J. Schwab et al. Real-time photoacoustic projection imaging using deep learning(2018).
[106] T. Karras et al. Progressive growing of GANS for improved quality, stability, and variation(2017).
[113] L. R. Medsker, L. Jain. Recurrent Neural Networks(2001).
[121] G. P. Luke et al. O-Net: a convolutional neural network for quantitative photoacoustic image segmentation and oximetry(2019).
[123] J. Gröhl et al. Estimation of blood oxygenation with learned spectral decoloring for quantitative photoacoustic imaging (LSD-qPAI)(2019).
[149] S. Ren et al. Faster R-CNN: towards real-time object detection with region proposal networks(2015).
[158] E. Moen et al. Deep learning for cellular image analysis. Nat. Methods, 16, 1233-1246(2019).
[159] S. Misra et al. Deep learning-based multimodal fusion network for segmentation and classification of breast cancers using B-mode and elastography ultrasound images. Bioeng. Transl. Med., e10480(2022).
[162] C. Szegedy et al. Going deeper with convolutions, 1-9(2015).
[168] C. Cortes, V. Vapnik. Support-vector networks. Mach. Learn., 20, 273-297(1995).
[173] M. Schwarz et al. Motion correction in optoacoustic mesoscopy. Sci. Rep., 7, 10386(2017).
[177] M. Jaderberg, K. Simonyan, A. Zisserman. Spatial transformer networks(2015).
[179] I. Gulrajani et al. Improved training of Wasserstein GANS(2017).
[188] P. Isola et al. Image-to-image translation with conditional adversarial networks, 1125-1134(2017).
[193] J.-Y. Zhu et al. Unpaired image-to-image translation using cycle-consistent adversarial networks, 2223-2232(2017).
[195] S. Bohndiek. Addressing photoacoustics standards. Nat. Photonics, 13, 298-298(2019).
[206] S. Choi et al. Synchrotron x-ray induced acoustic imaging. Sci. Rep., 11, 4047(2021).
Set citation alerts for the article
Please enter your email address



