Jianyu Yang, Hao Dong, Fulin Xing, Fen Hu, Leiting Pan, Jingjun Xu. Single-Molecule Localization Super-Resolution Microscopy and Its Applications[J]. Laser & Optoelectronics Progress, 2021, 58(12): 1200001
Search by keywords or author
- Laser & Optoelectronics Progress
- Vol. 58, Issue 12, 1200001 (2021)
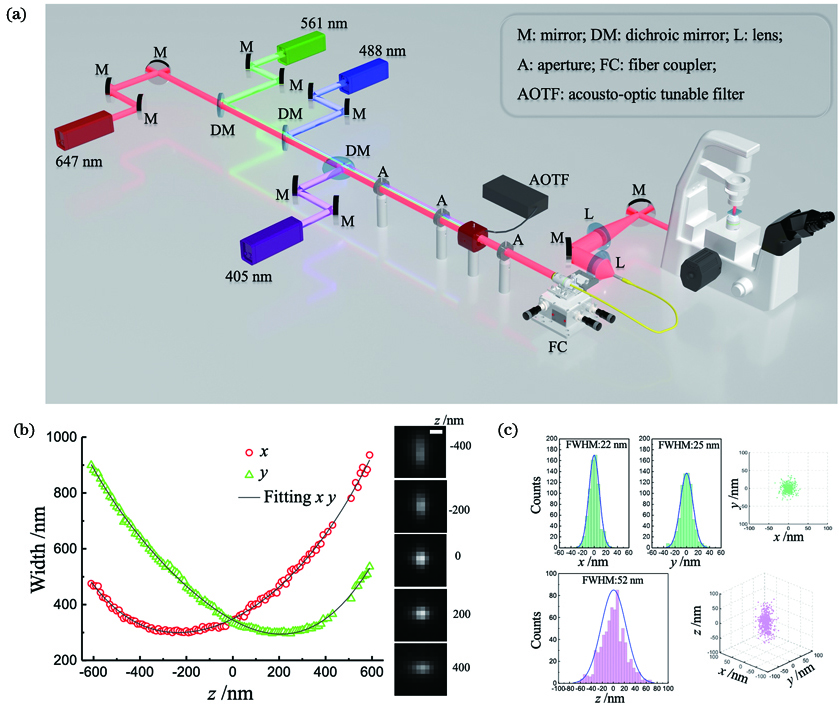
Fig. 1. Principle of 3D-STORM. (a) Optical path diagram of STORM; (b) PSF shape of a fluorophore at various z positions and calibration curves show x and y widths as functions of z (scale bar: 500 nm); (c) 3D localization distribution of single molecule fluorescence
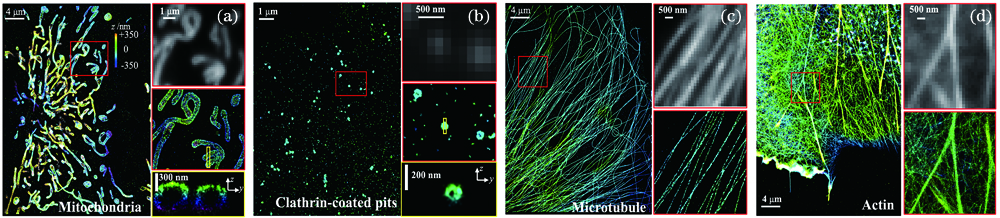
Fig. 2. STORM images of four subcellular structures from Cos7 cell. (a) Mitochondria; (b) clathrin-coated pits; (c) microtubules; (d) microfilaments
Fig. 3. Drift correction of STORM images of tubulin and tropomodulin in x and y directions. (a) STORM images of Cos7 cell microtubule before and after drift correction; (b) STORM images of erythrocyte tropomodulin before and after drift correction; (c) drift correction curves of microtubule images in x and y directions; (d) drift correction curves of erythrocyte tropomodulin in x and y directions
Fig. 4. Images of the same subcellular structures using different sample preparation methods. (a)(b) Images of mitochondria of Cos7 cell labeled with different primary antibodies; (c) images of microtubule of Cos7 cell fixed with methanol; (d) images of microtubule of Cos7 cell fixed with 3% paraformaldehyde+0.1% glutaraldehyde
Fig. 5. Improvement of SMLM lateral resolution based on algorithms or hardware modifications. (a) Improvement of STORM lateral resolution by dual objective fluorescence collection system[64]; (b) improvement of STORM lateral resolution by particle average algorithm[65]; (c) improvement of lateral resolution by MINFLUX based on algorithm combined hardware modification[14]; (d) improvement of lateral resolution by ROSE based on algorithm combined hardware modification [66]
Fig. 6. Improvement of SMLM axial resolution based on algorithms or hardware modifications. (a) Improvement of STORM axial resolution by dual objective fluorescence collection system[64]; (b) improvement of axial resolution by dual objective fluorescence interferometric system [71]; (c) improvement of DNA-PAINT axial resolution by developing new 3D fitting localization algorithm of PSF[72]; (d) improvement of axial resolution by ROSE-Z based on algorithm and hardware modification[73]
Fig. 7. Imaging of SMLM in large field of view (FOV). (a) Enlargement of the FOV of STORM to 100 μm×100 μm by FIFI[75]; (b) enlargement of the FOV of STORM to 221 μm×221 μm by multi-mode fiber[76]; (c) STORM imaging in large FOV of 500 μm×500 μm based on chip[77]
Fig. 8. Enlargement of SMLM axial range based on PSF engineering and modification of excitation light source. (a) Enlargement of axial range by self bending PSF engineering[81]; (b) enlargement of axial range by PSF engineering via tetrapod phase plate[63]; (c) enlargement of axial range by using oblique-plane illuminating system[85]
Fig. 9. Confocal/spectrum-SMLM correlative imaging. (a) Confocal-STORM correlative imaging of junctophilin, ryanodine receptors, and cell membrane[88]; (b) confocal-STORM correlative imaging of biocytin-filled axon terminals and cannabinoid receptor CB1[89]; (c) heterogeneity of cell membrane revealed by spectrally resolved PAINT[90]; (d) identification of surface adsorption layer of two-component mixture resolved by spectrally resolved PAINT[91]
Fig. 10. EM-SMLM correlative imaging. (a) Correlative imaging of mitochondria by EM and PALM (labeling TOM20 proteins) [93]; (b) correlative imaging of cytoskeletons and mitochondria by EM and STORM (labeling microtubules and mitochondria) [96]; (c) correlative imaging of FIB-SEM and SMLM for whole cells[98]
Fig. 11. AFM-SMLM correlative imaging. (a) AFM-STORM correlative imaging of Huntington protein aggregate[99]; (b) AFM-STORM correlative imaging of paxillin and lamellipodia extension in live cell[100]; (c) distributions of Na+/K+ATPase on membrane revealed by AFM-STORM correlative imaging[101]
Fig. 12. Principle and applications of Ripley’s K function. (a) Schematic diagram of Ripley’s K function[108]; (b) Ripley’s K function reveals the clustering of SIRPα in activated human macrophage[114]; (c) Ripley’s K function indicates cluster sizes of GLUT1 proteins on the apical/basal surface in Hela cells[115]
Fig. 13. Analysis results of pair correlation function. (a) Results of pair correlation function with random and clustered distributions[116]; (b) pair correlation function analysis results of CD47 in young and old mouse erythrocytes[120]
Fig. 14. Applications of auto/cross-correlation analysis. (a) One-dimensional cross-correlation analysis of actin, C-terminal of spectrin, N-terminal of spectrin, and adducin in axons[121]; (b) two-dimensional auto-correlation analysis of spectrin in the soma of neurons[122]; (c) across-correlation analysis of N-terminal of spectrin and cross-correlation analysis of C-terminal and N-terminal of spectrin in human erythrocytes [123]
Fig. 15. Principle and applications of DBSCAN and FOCAL. (a) Principle of DBSCAN[108]; (b) cluster analysis of tyrosine kinase in human and mouse platelets using DBSCAN[127]; (c) principle of FOCAL[128]; (d) cluster analysis of NPC in U2OS cells by FOCAL3D[129]
Fig. 16. Principle and applications of SR-Tesseler based on Voronoï diagram. (a) Principle of Voronoï diagram[131]; (b) automatic segmentation of the uniform (left) and non-uniform (right) simulated images by SR-Tesseler[131]; (c) segmentation of GluA1 SMLM image by SR-Tesseler[131]; (d) cluster analysis of ROP6 SMLM image by SR-Tesseler[132]; (e) cluster analysis of nuclear pore complex SMLM image by SR-Tesseler[133]
Fig. 17. Principle of DNA origami and identification of nanoclusters of membrane proteins. (a) Principle of DNA origami[139]; (b) simulation images of random membrane proteins with gradually increased labeling density[140]; (c) simulation images of clustered membrane proteins with gradually increased labeling density[140]; (d) quantitative analysis of nanoclusters (duty ratio η, cluster density ρ, and their normalized curves)[140]
Fig. 18. Delicate cytoskeletal ultrastructures revealed by STORM. (a) Membrane-associated periodic skeleton in nerve axons revealed by STORM[121]; (b) characteristics of membrane skeleton ultrastructure of erythrocyte under physiological conditions revealed by STORM[123]; (c) disassembly characteristics of intermediate filaments revealed by STORM[145]
Fig. 19. Applications of SMLM in visualizing subcellular structures, membrane proteins, and intracellular organelles. (a) PALM maps the fine structure of focal adhesions with axial dimension[148]; (b) STORM reconstructs the radial ninefold symmetry of centriole-containing complex[149]; (c) STORM reveals the structures of the ESCRT-III complex during abscission of the intercellular bridge connecting two dividing cells[150]; (d) nanoscale clustering of T cell receptor-mediated signaling complexs revealed by PALM[151]; (e) Nanoscale clustering of ryanodine receptors revealed by STORM and PAINT[155]; (f) 3D-STORM reveals the colocalization of purinosomes with mitochondria[159]
Fig. 20. Study on nanoscale diffusion rate of biomolecules in live cell by SMdM. (a) SMdM diffusivity map of free mEos3.2 in the cytoplasm of a live cell and correlated STORM image of the actin cytoskeleton[164]; (b) 3D-PAINT and SMdM images of live cell membranes [165]
|
Table 1. Comparison of different multi-color SMLMs
|
Table 2. Comparison of parameters of different methods to achieve large field SMLM imaging
|
Table 3. Comparison of different cluster analysis methods

Set citation alerts for the article
Please enter your email address



