- Advanced Photonics
- Vol. 4, Issue 6, 066001 (2022)
Abstract
Keywords
1 Introduction
Optical diffraction tomography (ODT) is an imaging technique for extracting the three-dimensional (3D) refractive index distribution of a sample, e.g., a biological cell using multiple two-dimensional (2D) images acquired at different illumination angles. The refractive index of the sample provides useful morphological information, making ODT an interesting approach for biological applications.1
In the last several years, many different iterative methods have been proposed to reconstruct accurate refractive indices from ill-posed measurements.5
PINNs have recently gotten intense research attention for solving different complex problems in physics.10,11 These networks use physics laws as the loss function instead of the data-driven loss functions. In conventional supervised deep learning, a large dataset of labeled examples is used for the training process: by comparing the known ground truth with the predictions from a deep multi-layer neural network, one can construct a loss function and tune the parameters of the network to solve complex physical problems. Different examples of these data-driven neural networks are proposed for optical applications such as resolution enhancement,12 imaging through multi-mode fibers,13,14 phase retrieval,15 ODT,16 and digital holography.17,18 In these networks, the knowledge acquired by the network strongly depends on the statistical information provided in the dataset, and training such a network requires access to a large dataset. In contrast, PINNs directly minimize the physical residual from the corresponding partial differential equation (PDE) that governs the problem instead of extrapolating physical laws after going through a large amount of examples. In the pioneering approach proposed by Lagaris et al.,19 the neural network maps independent variables, such as spatial and time coordinates, to a surrogate solution of a PDE. By applying the chain rule, e.g., through auto-differentiation integrated in many deep-learning packages, one can easily extract the derivatives of the output fields with respect to the input coordinates and consequently construct a physics-based loss.20 The correct prediction can be therefore retrieved by minimizing the loss with respect to the network weights. This approach has been used to solve nonlinear differential equations,21
Sign up for Advanced Photonics TOC. Get the latest issue of Advanced Photonics delivered right to you!Sign up now
Having the independent variables of PDE as the input of the neural network limits the use of PINNs when fast inference is required. For the example of optical scattering, the neural network should be trained for each refractive index distribution separately. A different idea was proposed recently in Ref. 27 to solve Maxwell’s equations for micro lenses with different permittivity distributions. The calculation of physical loss, in this case, is based on the finite difference scheme, and in contrast to the previous approach that is trained for a single example, this model proved to be well-suited for cases in which fast inference is required. However, such a PINN was only demonstrated to work for homogeneous 2D samples.
In this paper, we extend this idea for inhomogeneous and 3D cases and present a MaxwellNet that is able to solve different forward scattering problems, such as light scattering from biological cells. In the first part of the work, we train MaxwellNet for 2D digital phantoms and show how this pretrained network can be fine-tuned to predict light scattering from more complex and experimentally relevant samples, in our case, HCT-116 cells. We benchmark the performance of MaxwellNet in solving scattering problems for 2D and 3D objects. Next, we demonstrate that such PINN can be efficiently used to invert the scattering problem through an iterative scheme and improve the results of conventional ODT. We first demonstrate the reconstruction of the refractive index distribution from synthetic data and then we validate the technique with experimental measurements of scattering from polystyrene microspheres.
2 Methodology
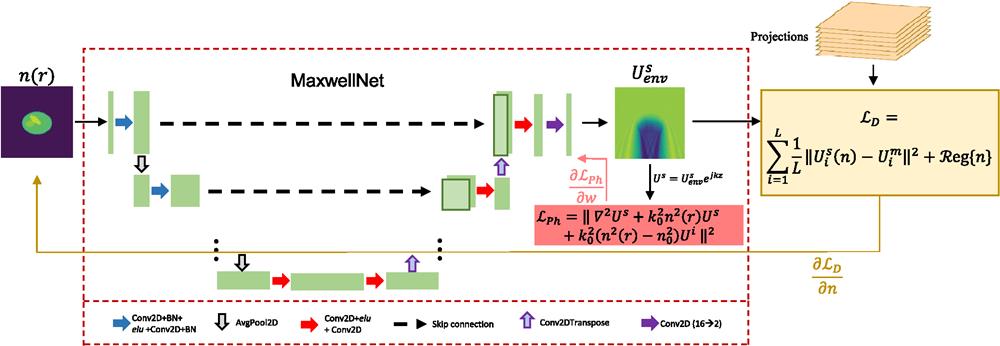
The main idea of our work, shown in Fig. 1, consists of two blocks. The first, MaxwellNet, is a neural network that takes as an input the refractive index distribution
![]()
Figure 1.Schematic description of MaxwellNet, with U-Net architecture, and its application for tomographic reconstruction. The input is a refractive index distribution and the output is the envelope of the scattered field. The output is modulated by the fast-oscillating term
2.1 Forward Model: MaxwellNet
In this section, we describe the implementation of a PINN that predicts the scattered field for a known input RI distribution. For the sake of simplicity, we first describe the method for the 2D case, but we will show the extension to 3D in the following. In this case, MaxwellNet takes as an input the RI distribution as a discrete array of shape
When we train MaxwellNet for a class of samples, it can accurately calculate the field for unseen samples from the same class. However, the key point to mention is that if we want to use MaxwellNet for a different set of RI distributions, we can fix some of the weights and adjust only a part of the network for the new dataset instead of re-starting the training from scratch. This process, referred to in the following as fine-tuning, is much faster than the original training of MaxwellNet. We will elaborate and discuss this interesting feature in Sec. 3.
It should be mentioned that we train MaxwellNet based on the Helmholtz equation with a scalar field approximation, as described in Eq. (1). The scalar approximation allows us to have a network with 2-channel output, representing the real and imaginary parts of the scalar field. We can also consider the full-vectorial Helmholtz equation where we need a larger network with 6-channel output to represent the real and imaginary parts of the three components of the field vector. However, the depolarization term can be neglected for samples with low refractive index gradients,31,32 allowing us to have a MaxwellNet with fewer parameters and the scalar Helmholtz equation as the loss function.
2.2 Optical Diffraction Tomography Using MaxwellNet
Once MaxwellNet has been trained on a class of RI distributions, it can be used to rapidly backpropagate reconstruction errors with an approach similar to learning tomography.6 Let us assume that we measure
Also, in this case, we use an Adam optimizer for updating the RI values. The regularizer in Eq. (3) consists of three parts: a total-variation (TV), a non-negativity, and physics-informed terms,
Importantly, we have to remark that MaxwellNet is trained for a specific dataset and accurately predicts the scattered field for RI distributions that are not too far from this set. To take into account this effect, we add the physics-informed loss to the regularizer. This further correction term helps to find RI values in a way that MaxwellNet can predict the scattered field for them correctly. In contrast to TV and non-negativity constraints that are used due to the ill-posedness of the ODT problem, the physics-informed regularizer is necessary in our methodology to ensure that the index distributions remain within the domain in which MaxwellNet has been trained.
The key advantages of using MaxwellNet with respect to other forward models are: differently from BPM, it can accurately calculate field scattering, considering reflection, multiple-scattering, or any other electromagnetic effects;5
3 Results and Discussion
3.1 MaxwellNet Results
In this section, we evaluate the performance of MaxwellNet for the prediction of the scattered field from RI structures such as biological cells. First, we check the performance on a 2D sample assuming that the system is invariant along the
In Figs. 2(a) and 2(b), we choose two random examples of the digital phantoms in the test set (which is not seen by the network during the training). For each test case, in the second and third rows, we present the prediction of the envelope of the scattered field by the network, and we compare it with the result achieved by the FEM using COMSOL Multiphysics 5.4. We can see a very small difference between the results of MaxwellNet and COMSOL, which we attribute to discretization errors. There are different schemes of discretization in two methods that can cause such differences. To quantitatively evaluate the performance of MaxwellNet, we define the relative error of MaxwellNet with respect to COMSOL as
![]()
Figure 2.Results of MaxwellNet and its comparison with COMSOL. (a) and (b) Two test cases from the digital phantom dataset and the prediction of the real and imaginary parts of the envelope of the scattered fields using MaxwellNet, COMSOL, and their difference. (c) Scattered field predictions from the network trained in (a) and (b) for the case of an experimentally measured RI of HCT-116 cancer cells and comparison with COMSOL. The difference between the two is no longer negligible. (d) Comparison between MaxwellNet and COMSOL after fine-tuning the former for a set of HCT-116 cells. MaxwellNet predictions reproduces much more accurate results after fine-tuning.
It should be noted that once MaxwellNet is trained, the scattered field calculation is much faster than numerical techniques such as FEM. We present a time comparison in Table 1. For the test phantoms in Fig. 2, it took 17 ms for MaxwellNet in comparison with 13 s for COMSOL, meaning three orders of magnitude acceleration.
| Dataset | 2D phantoms (training) | 2D HCT-116 (fine-tuning) | 3D phantoms (training) |
| Training details | 2700 samples 5000 epochs | 122 samples 600 epochs | 180 samples 5000 epochs |
| MaxwellNet training/fine-tuning | 30.5 h | 0.18 h | 15.5 h |
| MaxwellNet inference | 17 ms | 17 ms | 44.9 ms |
| COMSOL | 13 s | 13 s | 2472 s |
Table 1. Computation time comparison.
Furthermore, performing a physics-based instead of direct data-driven training holds promises for exploiting the advantages of transfer learning.33 Maxwell equations are general but having a neural network that predicts the scattered field for any class of RI distribution in milliseconds with a negligible physical loss is usually unfeasible. Most of the previous PINN studies for solving partial differential equations are trained for one example, and they will work for that specific example. In our case, the U-Net architecture proved to be expressive enough to predict the field for a class of samples. However, if we use MaxwellNet for inference on an RI distribution completely uncorrelated with the training set, the accuracy drops. To evaluate the MaxwellNet extrapolation capability, we considered the model trained on phantoms samples in Fig. 2 and use it for inference on HCT-116 cancer cells. The comparison between MaxwellNet and COMSOL is shown in Fig. 2(c). The input of the network is a 2D slice of the experimentally measured HCT-116 cell in the plane of the best focus. The discrepancy between MaxwellNet and COMSOL is due to the fact that the former does not see examples of such RI distributions during the training. As a result, if we require accurate results for a new set of samples with different features, we have to re-train MaxwellNet for the new dataset, which would take a long time as shown in Table 1. However, it turns out that learning a physical law, as Maxwell equations, even though on a finite dataset, is better suited than data-driven training for transfer learning on new batches. Indeed, we can use the pretrained MaxwellNet on digital phantoms and fine-tune some parts of the network for HCT cells achieving good convergence in a few epochs. In this example, we create a dataset of 136 RI distributions of HCT-116 cancer cells and divide them into the training and validation sets. Some examples of the HCT-116 refractive index dataset are shown in Appendix B. A wide range of cells with different shapes are included in the dataset. We have single cancer cells, such as shown in Fig. 2(c), examples of cells in the mitosis process, or examples with multiple cells. In this case, we freeze the weights of the encoder part and fine-tune the decoder with the new dataset. We can see in Fig. 2(d) that after this correction step, the calculated field is much more accurate. As can be seen in Table 1, the fine-tuning process is two orders of magnitude faster than a complete training from scratch.
The 2D case is helpful for demonstrating the method and rapidly evaluating the performances. Nevertheless, full 3D fields are required for many practical applications. We can straightforwardly recast MaxwellNet in 3D using arrays of size
The 3D version of MaxwellNet has
![]()
Figure 3.Results of 3D MaxwellNet and its comparison with COMSOL. The RI distribution is shown in (a). The real part of the envelope of the scattered field calculated by 3D MaxwellNet is shown in (b), calculated by COMSOL in (c), and their difference in (d). The imaginary part of the envelope of the scattered field calculated by 3D MaxwellNet, COMSOL, and their difference are presented in (e)–(g), respectively.
3.2 Tomographic Reconstruction Results
To show the ability of MaxwellNet to be used for different imaging applications, we implement an optimization task with MaxwellNet as the forward model for ODT, as explained in Sec. 2.2. In this example, we consider one of the digital phantoms in the test set of Fig. 2, and we use 2D MaxwellNet as the forward model to compute the 1D scattered field along the transverse direction
![]()
Figure 4.Tomographic reconstruction of RI using MaxwellNet. (a) The RI reconstruction was achieved by Rytov, MaxwellNet, and the ground truth. (b) 1D RI profile at
Next, we try a 3D digital phantom from the test set and we use the 3D MaxwellNet as the forward model in our tomographic reconstruction method. Since generating synthetic data with COMSOL is time-consuming for multiple angles, we create synthetic scattered fields from the phantom with the Lippmann–Schwinger equation. 9 Later, we will show an experimental example where we illuminate the sample with a circular illumination pattern with an angle
![]()
Figure 5.Tomographic RI reconstruction of 3D sample using MaxwellNet. The RI reconstruction is achieved by Rytov, MaxwellNet, learning tomography, and the ground truth in different rows at the
Additionally, we performed learning tomography6 for the synthetic measurements using 181 projections. The 3D tomographic reconstruction using learning tomography is shown in the third row of Fig. 5. In comparison with MaxwellNet, learning tomography has some elongated artifacts, which can be due to the fact that reflection is neglected in its forward model. However, the reconstruction with learning tomography is smoother in comparison with the reconstruction of MaxwellNet, which is slightly pixelated. This issue happens because the beam propagation method, as the forward model of learning tomography, is a smooth forward model with respect to the voxels of the refractive index distribution, which is not the case for a deep neural network such as MaxwellNet. However, the reconstructions are quantitatively comparable. If we assume the reconstruction error of
We also evaluated our methodology experimentally. We mentioned earlier that MaxwellNet takes care of reflection as a forward model, and therefore, our reconstruction technique can be used for samples with high contrast. In our experimental analysis, we try a polystyrene microsphere immersed in water, where we expect to have a
![]()
Figure 6.Tomographic RI reconstruction of a polystyrene microsphere immersed in water. The projections are measured with off-axis holography for different angles. The RI reconstructions achieved by Rytov, MaxwellNet, and learning tomography are presented at the
4 Conclusion
In summary, we proposed a PINN that rapidly calculates the scattered field from inhomogeneous RI distributions such as biological cells. Our network is trained by minimizing a loss function based on Maxwell equations. We showed that the network can be trained for a set of samples and could predict the scattered field for unseen examples that are in the same class. As our PINN is not a data-driven neural network, it can be trained for different examples in different conditions. Even though the network is not efficiently extrapolating to classes that are statistically very different from the training dataset, we showed that by freezing the encoder weights and fine-tuning the decoder branch, one can get a new predictive model in a few minutes. We believe that this can be further used for changing the wavelength, boundary condition, or other physical parameters.
We used our PINN as a forward model in an optimization loop to retrieve the RI distribution from the scattered fields achieved by illuminating the sample from different directions, known as ODT. This example shows the ability of MaxwellNet to be used as an accurate forward model in optimization loops for inverse design or inverse scattering problems.
5 Appendix A: Calculation of Physics-Informed Loss
During the training of MaxwellNet, we calculate at each epoch the loss function in Eq. (1) for the network output. To evaluate the Helmholtz equation residual, we should numerically compute the term
To minimize the discretization error, one can use a smaller pixel size,
6 Appendix B: Training and Fine-tuning of MaxwellNet
As mentioned in Sec. 3, we create a dataset of digital cell phantoms to train and validate MaxwellNet. The dataset for 2D MaxwellNet includes 3000 phantoms with elliptical shapes oriented in different directions. The size of these phantoms is in the range from 5 to
![]()
Figure 7.Training and fine-tuning of MaxwellNet. (a) Training (blue) and validation (orange) loss of MaxwellNet for the digital cell phantoms dataset. (b) Fine-tuning of the pretrained MaxwellNet for a dataset of HCT-116 cells for 1000 epochs. (c) Examples of the HCT-116 dataset.
We discussed in Sec. 3 using MaxwellNet that was trained for cell phantoms to predict the scattered field for real cells. A dataset of HCT-116 cancer cells is used for this purpose. The 3D refractive index of these cells is reconstructed using the Rytov approximation, with projections achieved with an experimental setup utilizing a spatial light modulator, as described in Ref. 8. Then, a 2D slice of the refractive index is chosen in the plane of best focus. A total number of 8 cells are used, and we rotated and shifted these cells to create a dataset of 136 inhomogeneous cells whose refractive index range is (1.33, 1.41). We use 122 of these images for training and 14 for validation. Some examples of the HCT-116 refractive index dataset are shown in Fig. 7(c). We freeze the encoder of MaxwellNet and fine-tune its decoder for this new dataset. The training and validation losses are shown in Fig. 7(b).
For 3D MaxwellNet, a dataset of 200 phantoms is created. These 3D phantoms have a spherical shape with some details inside them and the range of their diameter is 1.8 to
7 Appendix C: Experimental Setup for ODT
For ODT, we require complex scattered fields from multiple illumination angles. The off-axis holographic setup to accomplish that is shown in Fig. 8. It relies on an ytterbium-doped fiber laser at
![]()
Figure 8.Experimental setup for multiple illumination angle off-axis holography. HW: half-wave plate; P: polarizer; BS: beam splitter; L: lens; Obj: microscope objective; and M: mirror.
Amirhossein Saba is a PhD student in photonics at École Polytechnique Fédérale de Lausanne (EPFL), Lausanne, Switzerland. He received his BS degree in electrical engineering with a minor in physics from Sharif University of Technology, Tehran, Iran, in 2018. His current research interests include computational imaging, polarization-sensitive, and nonlinear imaging techniques.
Carlo Gigli received his MS degree in nanotechnologies for ICTs from Politecnico di Torino and Université Paris Diderot in 2017, and his PhD in physics at Université de Paris in 2021. During this period, his research interests included the design, fabrication, and characterization of dielectric resonators and metasurfaces for nonlinear optics. Since September 2021, he has been working as post-doc in the Optics Laboratory at EPFL, where his main activities focus on optical computing, physics informed neural network, and nonlinear tomography.
Ahmed B. Ayoub received his BS degree in electrical engineering from Alexandria University in Egypt in 2013. He received his MS degree in physics from the American University, Cairo, Egypt, in 2017. He is pursuing his PhD in electrical engineering at the École Polytechnique Fédérale de Lausanne, Lausanne, Switzerland. His current research interests include optical imaging and three-dimensional refractive index reconstructions for biological samples.
Demetri Psaltis received his BSc, MSc, and PhD degrees from Carnegie-Mellon University, Pittsburgh, Pennsylvania. He is a professor of optics and director of the Optics Laboratory at the École Polytechnique Fédérale de Lausanne (EPFL), Lausanne, Switzerland. In 1980, he joined the faculty at the California Institute of Technology, Pasadena, California. He moved to EPFL in 2006. His research interests include imaging, holography, biophotonics, nonlinear optics, and optofluidics. He has authored or coauthored over 400 publications in these areas. He is a fellow of the Optical Society of America, the European Optical Society, and SPIE. He was the recipient of the International Commission of Optics Prize, the Humboldt Award, the Leith Medal, and the Gabor Prize.
References
[1] W. Choi et al. Tomographic phase microscopy. Nat. Methods, 4, 717-719(2007).
[6] U. S. Kamilov et al. Learning approach to optical tomography. Optica, 2, 517-522(2015).
[10] G. E. Karniadakis et al. Physics-informed machine learning. Nat. Rev. Phys., 3, 422-440(2021).
[12] Y. Rivenson et al. Deep learning microscopy. Optica, 4, 1437-1443(2017).
[13] N. Borhani et al. Learning to see through multimode fibers. Optica, 5, 960-966(2018).
[22] S. M. H. Hashemi, D. Psaltis. Deep-learning PDEs with unlabeled data and hardwiring physics laws(2019).
[31] A. Ishimaru. Wave Propagation and Scattering in Random Media, 2(1978).
[32] A. Saba et al. Polarization-sensitive optical diffraction tomography. Optica, 8, 402-408(2021).
Set citation alerts for the article
Please enter your email address



