Author Affiliations
Institute of Optics Mechanics Electronics Technology and Application, East China Jiaotong University, Nanchang, Jiangxi 330013, Chinashow less
Fig. 1. Sample of camellia oleifera leaves. (a) Healthy camellia oleifera leaves; (b) infected anthracnose camellia oleifera leaves
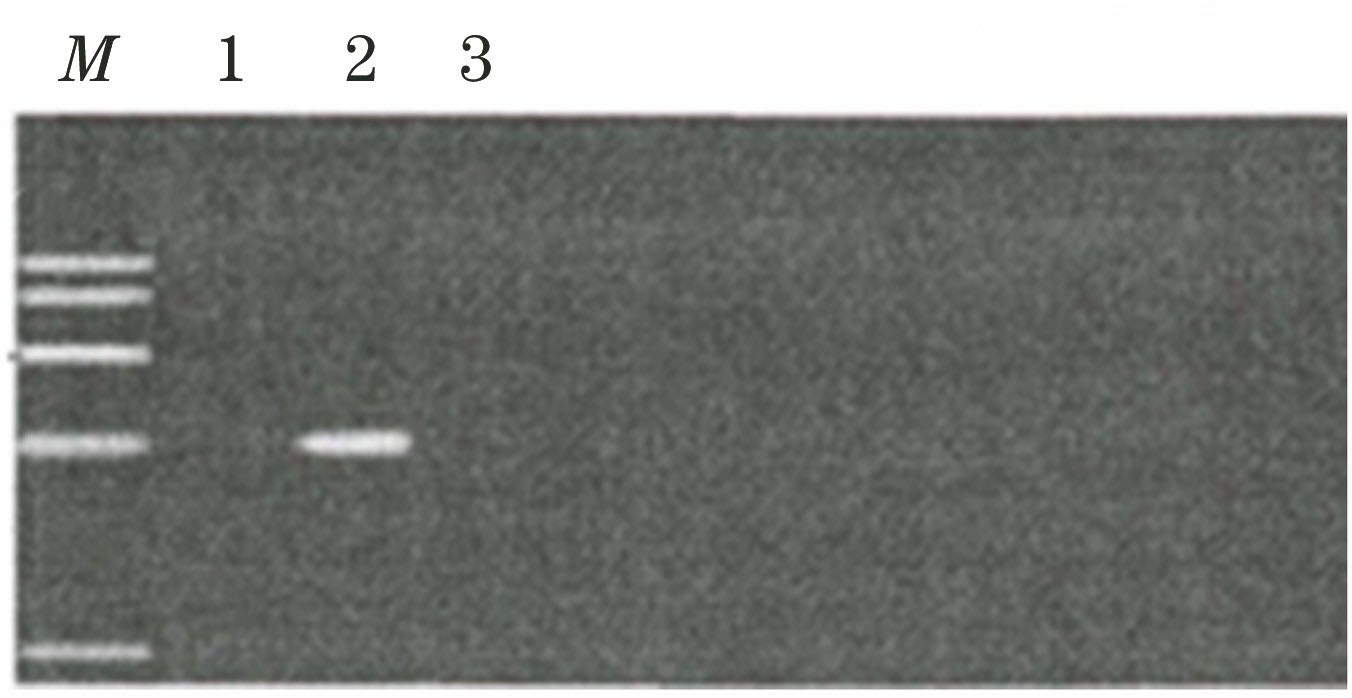
Fig. 2. PCR test results of camellia oleifera leaves
Fig. 3. Original spectrum of camellia oleifera leaves after interception
Fig. 4. Working curve of standard solution. (a) Healthy camellia oleifera leaves; (b) infected anthracnose camellia oleifera leaves
Fig. 5. Location of characteristic line of Mn element
Fig. 6. Comparison before and after data smoothing
Fig. 7. PLS model and prediction model of Mn element after 7-point data smoothing and first derivative de-noising. (a) PLS model; (b)prediction model
Fig. 8. Best sub-interval selected by the iPLS model. (a) Spectral graph of the 24th interval with RMSECV; (b) spectral graph corresponding to the 6th sub-interval
Fig. 9. iPLS modeling set scatter diagram
Fig. 10. iPLS prediction set scatter diagram
| Testcondition | Wavelength /nm | Lampcurrent /mA | Acetylene flowrate /(L·min-1) | Airflowrate /(L·min-1) | Slitwidth /nm |
|---|
| Parameter | 279.5 | 3 | 1.3 | 7.5 | 0.2 |
|
Table 1. Determination conditions of Mn element
| Sample | Category | Numberof samples | Rangevalue /(mg·mg-1) | Averagevalue /(mg·mg-1) |
|---|
| Healthy camelliaoleifera leaves | calibration set | 181 | 1.0600-4.2570 | 2.2919 | | prediction set | 59 | 1.4610-3.7240 | 2.2767 | | Camellia oleifera leaves with anthracnose | calibration set | 157 | 0.7990-3.3290 | 1.4476 | | prediction set | 51 | 1.0680-2.7880 | 1.3581 |
|
Table 2. Division of samples
| Spectral pretreatmentmethod | Evaluationindex | Beforesmoothing | 5 pointssmoothing | 7 pointssmoothing | 9 pointssmoothing |
|---|
| RC | 0.8461 | 0.8621 | 0.8956 | 0.8760 | | Original | RMSECV /(μg·mg-1) | 0.2575 | 0.2499 | 0.2204 | 0.2433 | | RP | 0.8195 | 0.8212 | 0.8540 | 0.8315 | | RMSEP /(μg·mg-1) | 0.2769 | 0.2658 | 0.2548 | 0.2591 | | RC | 0.8534 | 0.8562 | 0.8648 | 0.8605 | | Denoising | RMSECV /(μg·mg-1) | 0.2434 | 0.2375 | 0.2403 | 0.2482 | | RP | 0.8162 | 0.8196 | 0.8329 | 0.8142 | | RMSEP /(μg·mg-1) | 0.2553 | 0.2469 | 0.2373 | 0.2499 | | RC | 0.8345 | 0.8440 | 0.8840 | 0.8396 | | Baseline correction | RMSECV /(μg·mg-1) | 0.2875 | 0.2570 | 0.1770 | 0.2764 | | RP /(μg·mg-1) | 0.8010 | 0.8219 | 0.8333 | 0.8019 | | RMSEP /(μg·mg-1) | 0.3169 | 0.2722 | 0.2524 | 0.2911 | | RC | 0.8584 | 0.8947 | 0.9025 | 0.8892 | | First derivativede-noising | RMSECV /(μg·mg-1) | 0.2414 | 0.2206 | 0.2192 | 0.2227 | | RP | 0.8215 | 0.8519 | 0.8882 | 0.8352 | | RMSEP /(μg·mg-1) | 0.2692 | 0.2474 | 0.2356 | 0.2339 | | RC | 0.8523 | 0.8521 | 0.8507 | 0.8619 | | Second derivativede-noising | RMSECV /(μg·mg-1) | 0.2528 | 0.2532 | 0.2569 | 0.2354 | | RP | 0.8190 | 0.8150 | 0.8377 | 0.8323 | | RMSEP /(μg·mg-1) | 0.2784 | 0.2791 | 0.2774 | 0.2802 | | RC | 0.8323 | 0.8389 | 0.8429 | 0.8413 | | Normalization | RMSECV /(μg·mg-1) | 0.2829 | 0.2705 | 0.2711 | 0.2514 | | RP | 0.8189 | 0.8106 | 0.8254 | 0.8177 | | RMSEP /(μg·mg-1) | 0.3062 | 0.3098 | 0.2955 | 0.3016 |
|
Table 3. PLS model results of smoothed data processed by different preprocessing methods
| Interval number | Optimum principal component | RMSECV /(μg·mg-1) | R | Optimum interval |
|---|
| 1 | 7 | 0.2770 | 0.8304 | 1 | | 2 | 9 | 0.2580 | 0.8548 | 1 | | 3 | 10 | 0.2360 | 0.8807 | 1 | | 4 | 12 | 0.2200 | 0.8966 | 1 | | 5 | 6 | 0.2510 | 0.8632 | 2 | | 6 | 6 | 0.2490 | 0.8660 | 2 | | 7 | 6 | 0.2560 | 0.8577 | 2 | | 8 | 6 | 0.2550 | 0.8589 | 2 | | 9 | 7 | 0.2330 | 0.8843 | 3 | | 10 | 8 | 0.2350 | 0.8832 | 3 | | 11 | 7 | 0.2380 | 0.8797 | 3 | | 12 | 7 | 0.2400 | 0.8769 | 3 | | 13 | 7 | 0.2200 | 0.8966 | 4 | | 14 | 6 | 0.2260 | 0.8918 | 4 | | 15 | 7 | 0.2240 | 0.8931 | 4 | | 16 | 7 | 0.2270 | 0.8899 | 4 | | 17 | 6 | 0.2220 | 0.8945 | 5 | | 18 | 6 | 0.2210 | 0.8961 | 5 | | 19 | 7 | 0.2170 | 0.8999 | 5 | | 20 | 8 | 0.2200 | 0.8979 | 5 | | 21 | 6 | 0.2350 | 0.8816 | 6 | | 22 | 8 | 0.2160 | 0.9016 | 6 | | 23 | 8 | 0.2160 | 0.9013 | 6 | | 24 | 8 | 0.2090 | 0.9076 | 6 | | 25 | 6 | 0.2280 | 0.8889 | 7 | | 26 | 7 | 0.2100 | 0.9068 | 7 | | 27 | 6 | 0.2140 | 0.9027 | 7 | | 28 | 6 | 0.2090 | 0.9067 | 7 | | 29 | 8 | 0.2700 | 0.8414 | 7 | | 30 | 8 | 0.2190 | 0.8996 | 8 |
|
Table 4. iPLS modeling analysis results