Yiming Liu, Zhiyong Xiao. Automatic Segmentation Algorithm of Liver Tumor Based on Feature Fusion[J]. Laser & Optoelectronics Progress, 2021, 58(14): 1417001
Search by keywords or author
- Laser & Optoelectronics Progress
- Vol. 58, Issue 14, 1417001 (2021)
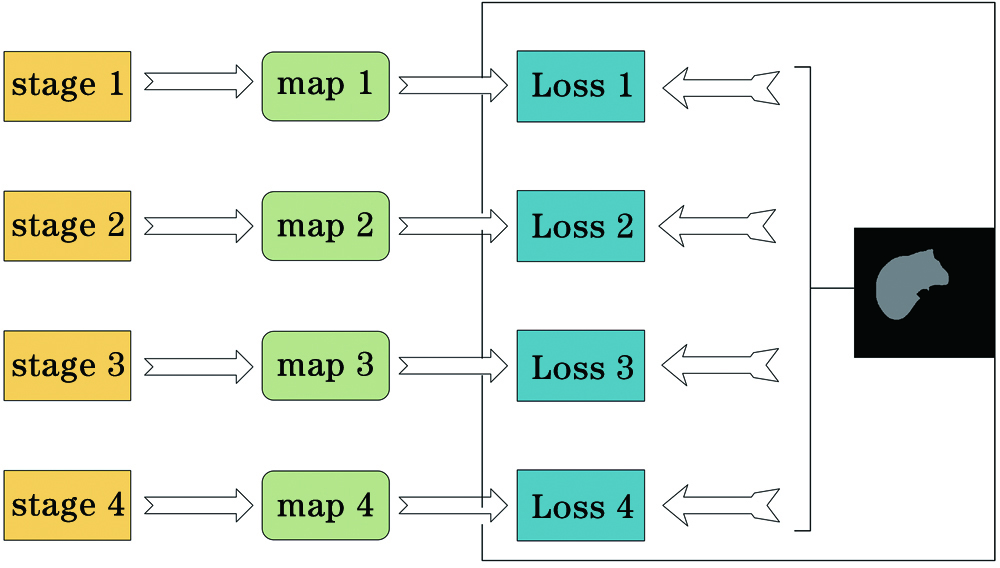
Fig. 1. Structure of deeply-supervised net
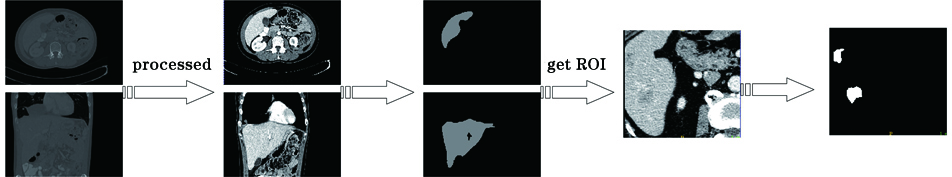
Fig. 2. Flow chart of our experiment
Fig. 3. Network structure proposed in this paper
Fig. 4. Attention module diagrams. (a) Channel attention module; (b) spatial attention module
Fig. 5. Comparison before and after pretreatment. (a) Transverse plane; (b) sagittal plane; (c) coronal plane; (d) HU distribution before pretreatment; (e) HU distribution after pretreatment
Fig. 6. Segmentation results of different methods. (a) Raw image; (b) Ground truth; (c) proposed method; (d) U-Net; (e) U-Net+deeply-supervised net; (f) U-Net+deeply-supervised net+spatial attention
Fig. 7. Comparison between proposed method and One-stage
|
Table 1. Segmentation results at different input sizes
| |||||||||||||||||||||||||||||||||||||||||
Table 2. Comparison of segmentation results of various network structures
| |||||||||||||||||||||||||||
Table 3. Comparison between proposed method and One-stage
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Table 4. Comparison of different segmentation methods

Set citation alerts for the article
Please enter your email address



