Zhushanying Zhang, Hanwen Gu, Kaiwen Xie, He Jiang, Qinlan Xie, Jiming Sa. Pretreatment and Combined Method Based on Near Infrared Spectroscopy[J]. Laser & Optoelectronics Progress, 2021, 58(16): 1617001
Search by keywords or author
- Laser & Optoelectronics Progress
- Vol. 58, Issue 16, 1617001 (2021)

Fig. 1. Physical image of the hemoglobin mimetic solution
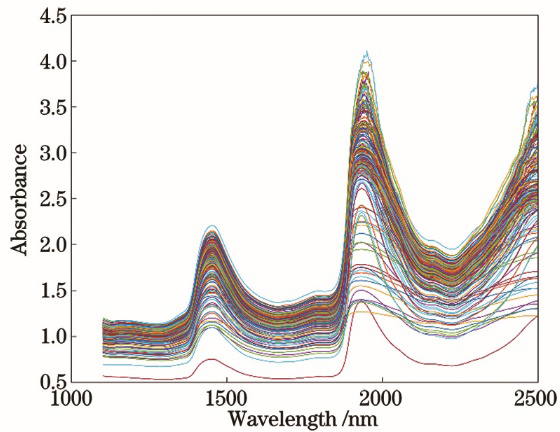
Fig. 2. Raw spectra of the blood samples
Fig. 3. Spectra of blood after removing singular samples
Fig. 4. NIR obtained by single pretreatment method. (a) Mean centering; (b) scaling; (c) normalization; (d) first derivative; (e) second derivative; (f) MSC; (g) SNV; (h) MWA smoothing; (i) SG smoothing
Fig. 5. Optimization of MWA parameters of blood samples
Fig. 6. RMSECV obtained by different pretreatment methods(blood)
Fig. 7. Raw spectra of the phantom solution
Fig. 8. Spectra of the phantom solution after removing singular samples
Fig. 9. Spectra of the phantom solution obtained by a single pretreatment method. (a) Mean centering; (b) scaling; (c) normalization; (d) first derivative; (e) second derivative; (f) MSC; (g) SNV; (h) MWA smoothing; (i) SG smoothing
Fig. 10. RMSECV obtained by different pretreatment methods (phantom solution)
|
Table 1. Raw material of the hemoglobin mimetic solution
|
Table 2. Hemoglobin contents of different data sets
|
Table 3. Classification of pretreatment methods
|
Table 4. Combination of pretreatment methods (blood)
|
Table 5. Combination of pretreatment methods (phantom solution)

Set citation alerts for the article
Please enter your email address



