Xusheng Li, Donghua Chen, Saisai Liu, Naiming Zhang, Hu Li. Tree-Species Identification of Multisource Remote-Sensing Data using Improved 3D-CNN[J]. Laser & Optoelectronics Progress, 2020, 57(24): 242804
Search by keywords or author
- Laser & Optoelectronics Progress
- Vol. 57, Issue 24, 242804 (2020)
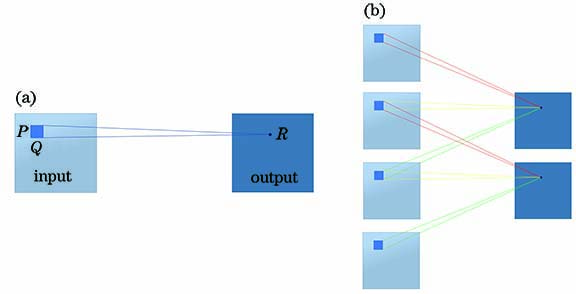
Fig. 1. Principle comparison between 3D-CNN and 2D-CNN. (a) 2D-CNN; (b) 3D-CNN
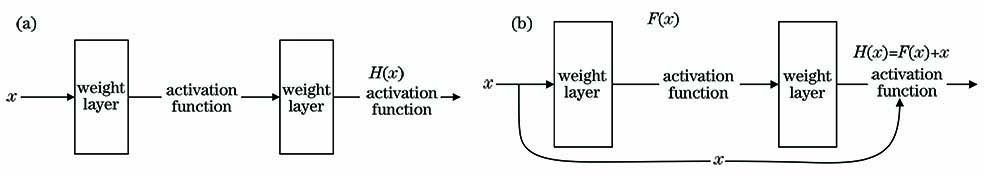
Fig. 2. Diagram of network structure. (a) Flat network; (b) ResNet
Fig. 3. Structure of 3D-RCNN
Fig. 4. Distribution of study area and samples
Fig. 5. Correlation matrix of characteristic factors. (a) Spectral feature; (b) texture feature; (c) vegetation index feature
Fig. 6. Sample expansion by inner ring rotation
Fig. 7. Influence of number of convolution units on test time and overall accuracy
Fig. 8. Influence of step size on test time and overall accuracy
Fig. 9. Tree species distribution. (a) Forest inventory; (b) algorithm extraction
|
Table 1. Basic information of remote-sensing image data
|
Table 2. Number of samples
|
Table 3. Influence of input pixel size on operation time and overall accuracy
|
Table 4. Influence of convolution kernel size on operation time and overall accuracy
|
Table 5. Influence of learning rate on convergence rate and overall accuracy
|
Table 6. Classification accuracy evaluation matrix of each algorithm
|
Table 7. Accuracy verification of tree species area

Set citation alerts for the article
Please enter your email address



