Hongyang Ruan, Zhilan Chen, Yingsheng Cheng, Kai Yang. Detection of Pulmonary Nodules Based on C-3D Deformable Convolutional Neural Network Model[J]. Laser & Optoelectronics Progress, 2020, 57(4): 041013
Search by keywords or author
- Laser & Optoelectronics Progress
- Vol. 57, Issue 4, 041013 (2020)
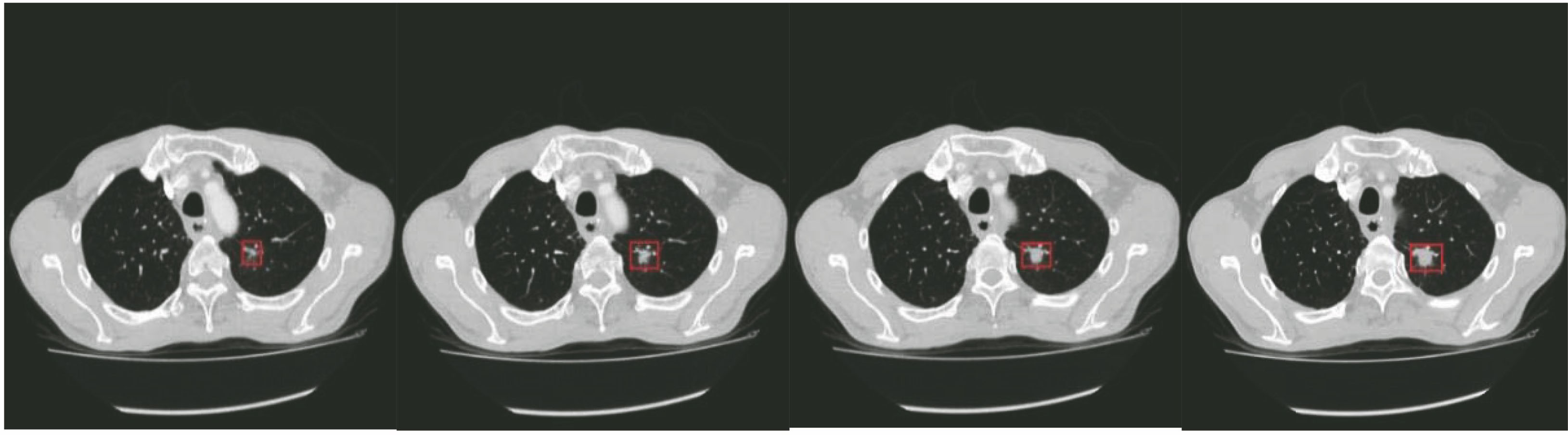
Fig. 1. CT images of lung
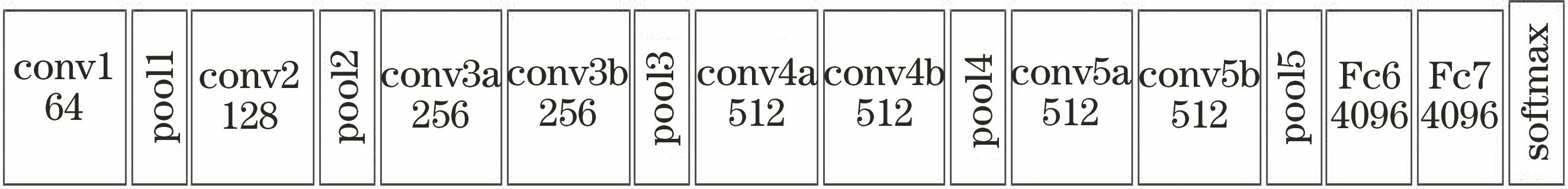

Fig. 2. Structure of C-3D convolutional neural network model
Fig. 3. Structures of 3D deformable convolution and pooling. (a) 3D deformable convolution; (b) 3D deformable pooling
Fig. 4. Structure of C-3D deformable convolutional neural network model
Fig. 5. Classification accuracy for different learning rates and optimization functions. (a) Experimental comparison of learning rate; (b) experimental comparison of optimization function
Fig. 6. ROC curves and PRC curves of different models. (a) ROC curves; (b) PRC curves
Fig. 7. Boxes of deformable convolution layers with different numbers of features. (a) Box of AUC; (b) box of F1; (c) box of P; (d) box of R
Fig. 8. Visualization results of convolution window pixel sampling. (a) Original labeled lung images; (b) original C-3D convolutional neural network; (c) improved C-3D convolutional neural network
|
Table 1. Setting of parameters of convolution layer and pooling layer in C-3D deformable convolutional neural network
|
Table 2. Dataset information
|
Table 3. Comparison of results of different convolutional neural network models
|
Table 4. Results comparison of different output features in C-3D deformable convolutional neural network
|
Table 5. Comparison of different convolutional neural networks

Set citation alerts for the article
Please enter your email address



