Weisong Zhao, Yuanyuan Huang, Zhenqian Han, Liying Qu, Haoyu Li, Liangyi Chen. Deconvolution in Super-Resolution Fluorescence Microscopy (Invited)[J]. Chinese Journal of Lasers, 2024, 51(1): 0107002
Search by keywords or author
- Chinese Journal of Lasers
- Vol. 51, Issue 1, 0107002 (2024)
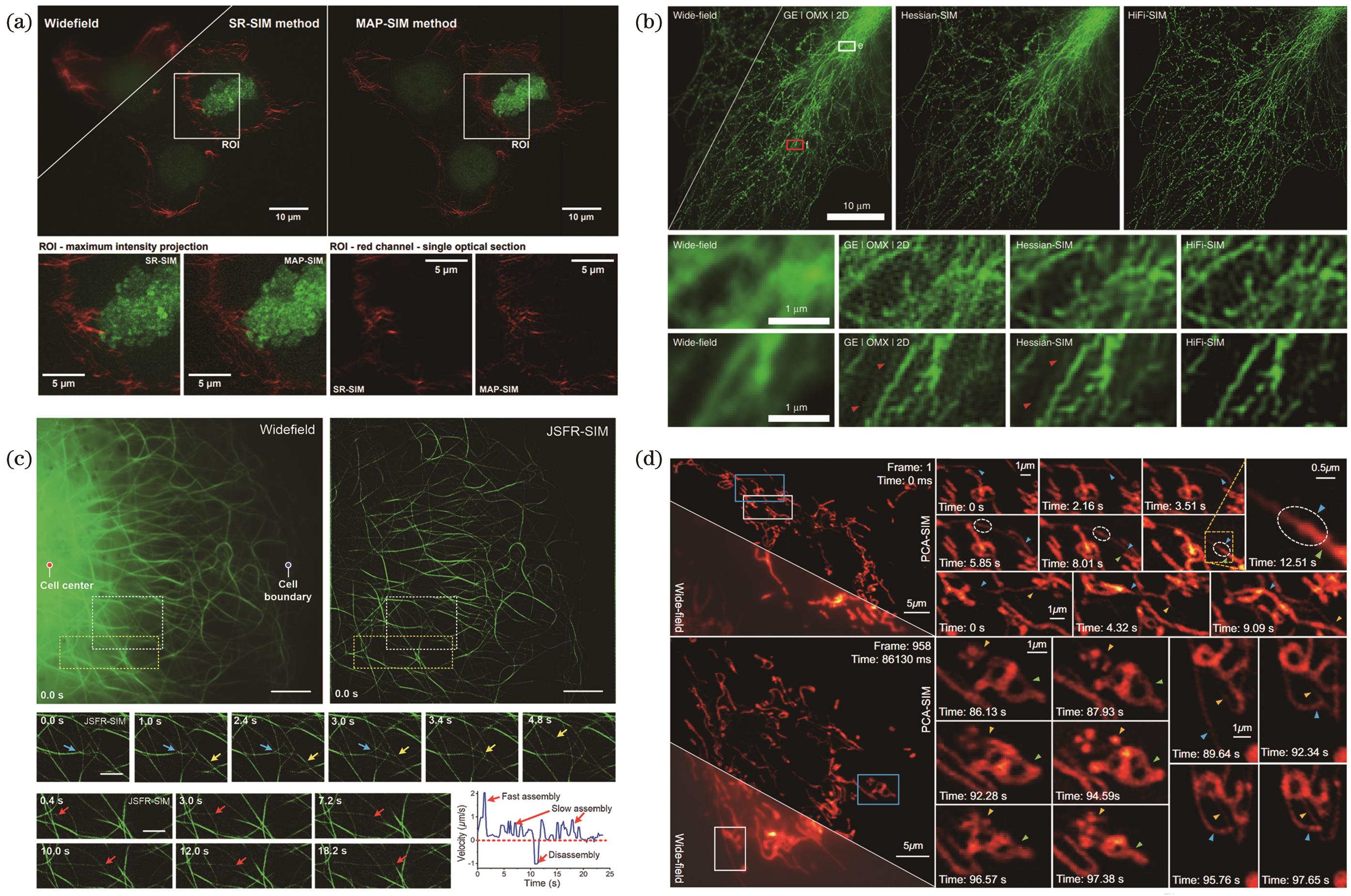
![Super-resolution imaging results based on deconvolution algorithms in structured illumination microscopy. (a) Dual-color three-dimensional super-resolution images of actin (magenta) and chromatin (green) based on MAP-SIM[33]; (b) HiFi-SIM reconstruction of microtubules in COS-7 cells[35]; (c) visualization of microtubule dynamics based on JSFR-SIM[36]; (d) super-resolution reconstruction of dynamic processes in COS-7 mitochondria based on PCA-SIM[37]](/richHtml/zgjg/2024/51/1/0107002/img_01.jpg)
Fig. 1. Super-resolution imaging results based on deconvolution algorithms in structured illumination microscopy. (a) Dual-color three-dimensional super-resolution images of actin (magenta) and chromatin (green) based on MAP-SIM[33]; (b) HiFi-SIM reconstruction of microtubules in COS-7 cells[35]; (c) visualization of microtubule dynamics based on JSFR-SIM[36]; (d) super-resolution reconstruction of dynamic processes in COS-7 mitochondria based on PCA-SIM[37]
![Super-resolution imaging results based on regularization deconvolution algorithms in structured illumination microscopy. (a) Dynamics of mitochondrial cristae structure in live cells based on Hessian-SIM[41]; (b) sorSIM reconstruction of microtubule-stained images in Vero cells[42]; (c) Sparse-SIM reconstruction of both inner and outer mitochondrial membranes in live COS-7 cells[8]; (d) super-fast imaging of fusion pores in INS-1 cells based on Sparse-SIM[8]; (e) super-resolution reconstruction results of Sparse-ExM[8]; (f) nulti-color 3D-SIM reconstruction of TV-FISTA-SIM (from left to right: outer mitochondrial membrane, microtubule protein, and actin imaging)45]](/richHtml/zgjg/2024/51/1/0107002/img_02.jpg)
Fig. 2. Super-resolution imaging results based on regularization deconvolution algorithms in structured illumination microscopy. (a) Dynamics of mitochondrial cristae structure in live cells based on Hessian-SIM[41]; (b) sorSIM reconstruction of microtubule-stained images in Vero cells[42]; (c) Sparse-SIM reconstruction of both inner and outer mitochondrial membranes in live COS-7 cells[8]; (d) super-fast imaging of fusion pores in INS-1 cells based on Sparse-SIM[8]; (e) super-resolution reconstruction results of Sparse-ExM[8]; (f) nulti-color 3D-SIM reconstruction of TV-FISTA-SIM (from left to right: outer mitochondrial membrane, microtubule protein, and actin imaging)45]
Fig. 3. Super-resolution imaging results based on deconvolution algorithms in improved structured illumination microscopy. (a) Imaging results of the wave-like protein network in fixed HUVEC cells based on RIM[48]; (b) three-dimensional imaging results based on RIM[48]; (c) mitochondrial fission and fusion processes mediated by the endoplasmic reticulum (magenta) based on GI-SIM[50]
Fig. 4. Imaging results based on deconvolution algorithms in image scanning microscopy. (a) Principle of MSIM image reconstruction and reconstruction of microtubules (green) and mitochondrial outer membrane (magenta)[52]; (b) principle of CSD-ISM image reconstruction and the resulting reconstruction[53]; (c) application of NNLS deconvolution to a test sample with 70 nm resolution[56]; (d) generation of ISM and Q-ISM images and reconstruction using the Joint Sparse Recovery (JSR) algorithm[60]; (e) Sparse SD-SIM imaging results of a four-color live cell, with lysosomes (yellow), mitochondrial matrix (green), cell nucleus (blue), and microtubule proteins (red)[8]; (f) Sparse SD-SIM imaging results of the endoplasmic reticulum in live cells[8]
Fig. 5. Stimulated emission depletion microscopy imaging results based on deconvolution algorithms. (a) RL-STED reconstruction results[63]; (b) smart RESOLFT reconstruction[64]; (c) Sparse-STED reconstruction of the nuclear pore protein[8]; (d) cross-validation of dual-color confocal microscopy and stimulated emission depletion microscopy, highlighting the mitochondrial outer membrane (magenta) and microtubule proteins (green)[8]
Fig. 6. Super-resolution optical fluctuation imaging results based on deconvolution algorithms. (a) Workflow of classical super-resolution optical fluctuation imaging techniques[66]; (b) Fourier SOFI resolution of the microtubule protein network in fibroblast cells[67]; (c) 3B analysis of focal adhesion proteins in fixed cells[70]; (d) SIMBA imaging of grid protein clusters (ccp) in live HeLa cells[71]; (e) multi-plane three-dimensional SOFI of live HeLa cells expressing vimentin-Dreiklang[73]
Fig. 7. Super-resolution optical fluctuation imaging results based on deconvolution algorithms. (a) SRRF applied to wide-field illumination microscopy[78]; (b) imaging of the endoplasmic reticulum in live Jurkat cells using eSRRF combined with lattice light-sheet microscopy[79]; (c) sparse deconvolution imaging of the mitochondrial outer membrane in HEK293-T cells[74]; (d) imaging of actin in live cells based on SOFI2.0[84]; (e) high-resolution four-dimensional imaging of live cell mitochondrial outer membrane based on SACD[87]; (f) high-throughput imaging of cell microtubules based on SACD[87]
Set citation alerts for the article
Please enter your email address



