Lihao Lin, Jianbing Yi, Feng Cao, Wangsheng Fang. Non-Rigid Registration Algorithm of Lung Computed Tomography Image Based on Multi-Scale Parallel Fully Convolutional Neural Network[J]. Laser & Optoelectronics Progress, 2022, 59(16): 1617004
Search by keywords or author
- Laser & Optoelectronics Progress
- Vol. 59, Issue 16, 1617004 (2022)
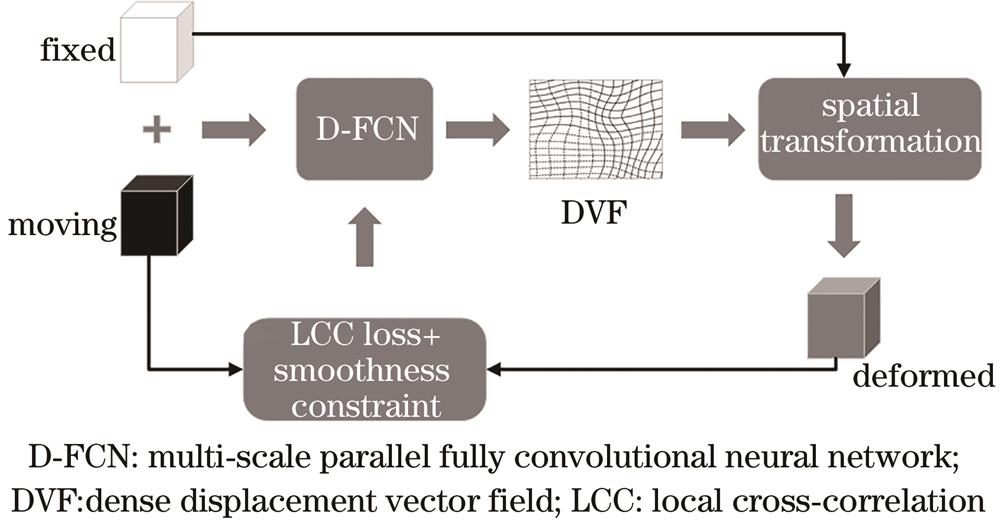
Fig. 1. Flow chart of D-FCN lung image registration algorithm
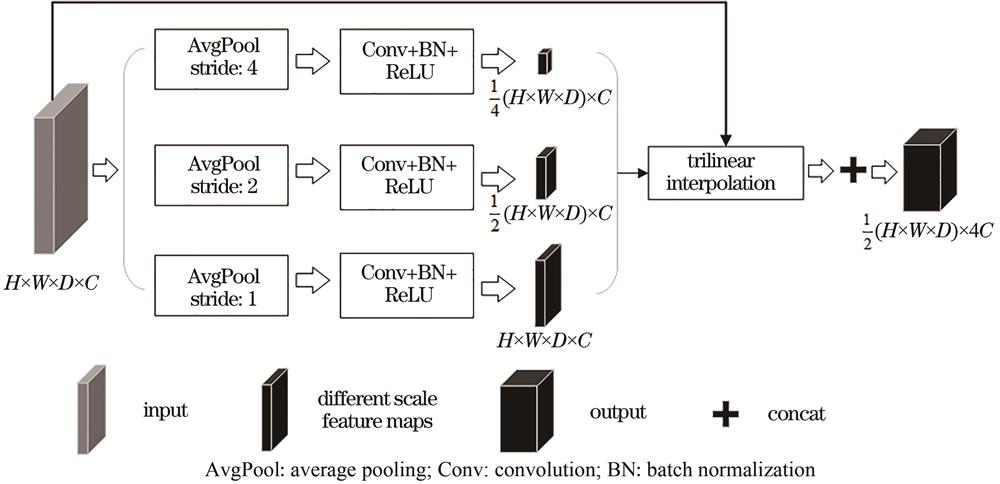
Fig. 2. Multi-scale parallel down-sampling module
Fig. 3. Lung feature maps at the same resolution and different scales. (a) Original image; (b) pooling core is 1; (c) pooling core is 2; (d) pooling core is 4
Fig. 4. Schematic diagram of dilated convolution. (a) Dilated factor is 1; (b) dilated factor is 2; (c) dilated factor is 3; (d) receptive fields with dilated factor of 1; (e) receptive fields with dilated factor of 2; (f) receptive fields with dilated factor of 3
Fig. 5. Pyramid dilated convolution module
Fig. 6. Different feature maps at 1/8 resolution after three down sampling. (a) Large deformation feature map; (b) small deformation feature map
Fig. 7. Adaptive channel attention module
Fig. 8. Structure diagram of dilated FCN
Fig. 9. Original lung CT image and images under noise attack (axial surface). (a) Original image; (b) image with Gaussian noise; (c) image with salt and pepper noise
Fig. 10. Registration results of case1 in DIR-lab dataset (coronal plane). (a) Fixed image; (b) moving image; (c) deformed image; (d) difference before registration; (e) difference after registration
Fig. 11. Box plot of target registration errors obtained by proposed algorithm from maximum inhalation to maximum exhalation in 10 cases in DIR-lab dataset
Fig. 12. Registration results of case1 in Creatis dataset (coronal plane). (a) Fixed image; (b) moving image; (c) deformed image; (d) difference before registration; (e) difference after registration
Fig. 13. Box plot of target registration errors obtained by proposed algorithm from maximum inhalation to maximum exhalation in 6 cases of Creatis dataset
|
Table 1. Target registration errors (standard deviations) of 300 expert points in DIR-lab dataset for each improvement step of proposed algorithm
|
Table 2. Target registration errors (standard deviations) of 300 expert points in DIR-lab dataset with different dilated factors in module
|
Table 3. Target registration errors (standard deviations) of 300 expert points in DIR-lab dataset of each registration algorithm
|
Table 4. Target registration errors (standard deviations) of 100 expert points in Creatis dataset of each registration algorithm
|
Table 5. Running time of each algorithm to register a set of lung CT images

Set citation alerts for the article
Please enter your email address



