Author Affiliations
11. School of Energy and Power Engineering, Jiangsu University of Science and Technology, Zhenjiang 212003, China22. National Research Ogarev Mordovia State University, Saransk 430005, Russia33. School of Materials Science and Engineering, Xi’an University of Science and Technology, Xi’an 710054, China44. State Key Laboratory of High Performance Ceramics and Superfine Microstructure, Shanghai Institute of Ceramics, Chinese Academy of Sciences, Shanghai 200050, Chinashow less
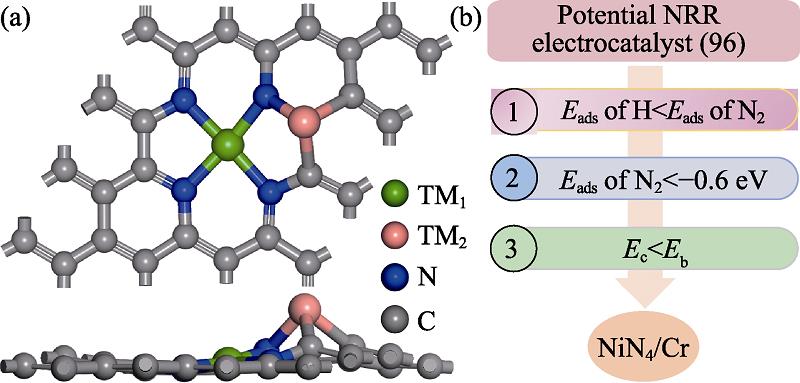
1. (a) Atomic structure of TM1N4/TM2 and (b) screening criterion for TM1N4/TM2 combination
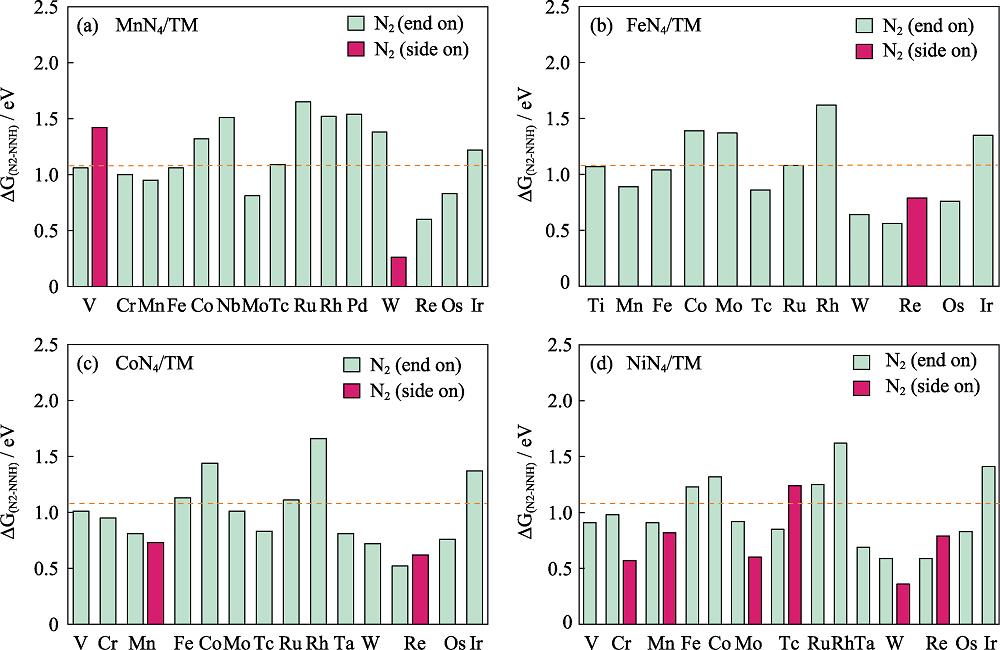
2. Gibbs free energy difference between N2 and NNH with the orange line at 1.08 eV
Colorful figures are available on website
3. Schematic reaction mechanismsColorful figure is available on website
4. Free energy diagrams and the corresponding configuration of the NRR intermediates on NiN4/Cr
NRR mechanisms are (a) distal, (b) alternating and (c) enzymatic
5. (a-c) Charge variation of the three moieties along the optimal pathway and (d) N-N bond length change in NRR along preferred pathwayMoieties 1, 2, 3 represent the graphene substrate, active center, and NRR intermediates, respectively
S1. Comparison of binding energy and bulk cohesive energy of the selected complexes
S2. Free energy diagrams and the corresponding configuration of the NRR intermediates on NiN4/Mo
NRR mechanisms are (a) distal, (b) alternating, and (c) enzymatic, respectively
S3. Free energy diagrams and the corresponding configuration of the NRR intermediates on NiN4/Ta
NRR mechanisms are (a) distal, (b) alternating, and (c) enzymatic, respectively
S4. Free energy diagrams and the corresponding configuration of the NRR intermediates on Cr embedded nitrogen functionalized grapheneNRR mechanisms are (a) distal, (b) alternating, and (c) enzymatic, respectively
S5. Free energy diagrams and the corresponding configuration of the NRR intermediates on Mo embedded nitrogen functionalized grapheneNRR mechanisms are (a) distal, (b) alternating, and (c) enzymatic, respectively
S6. Free energy diagrams and the corresponding configuration of the NRR intermediates on Ta embedded nitrogen functionalized grapheneNRR mechanisms are (a) distal, (b) alternating, and (c) enzymatic, respectively
S7. Free energy profiles of N2-to-NH3 conversion on the N vacancy
S8. Atomic configurations of the hydrogen adsorption on the nitrogen embedded in graphene after geometry optimization
| 3d | Sc | Ti | V | Cr | Mn | Fe | Co | Ni |
|---|
| Eads(TM2) N2 end-on
| -0.22 | -0.36 | -0.62 | -0.72 | -1.02 | -1.07 | -0.89 | -0.59 | | Eads(TM2) N2 side-on
| 0.12 | -0.02 | -1.17 | -0.35 | -0.59 | -0.51 | -0.33 | -0.19 | | Eads(TM2) H
| 0.75 | 0.20 | -0.18 | -0.14 | -0.19 | -0.20 | -0.22 | -0.38 | | 4d | Y | Zr | Nb | Mo | Tc | Ru | Rh | Pd | | Eads(TM2) N2 end-on
| -0.14 | -0.22 | -1.05 | -0.70 | -0.73 | -0.99 | -0.73 | -1.30 | | Eads(TM2) N2 side-on
| -0.13 | 0.11 | -0.42 | -0.43 | -0.47 | -0.44 | -0.25 | -0.96 | | Eads(TM2) H
| 0.78 | 0.25 | -0.87 | -0.38 | 0.51 | -0.11 | -0.33 | -1.06 | | 5d | Lu | Hf | Ta | W | Re | Os | Ir | Pt | | Eads(TM2) N2 end-on
| -0.21 | -0.35 | -0.60 | -1.57 | -1.23 | -1.30 | -1.08 | -0.52 | | Eads(TM2) N2 side-on
| 0.07 | 0.02 | -0.32 | -1.48 | -0.88 | -0.68 | -0.44 | -0.23 | | Eads(TM2) H
| 0.65 | -0.01 | -1.33 | -0.92 | -0.88 | -0.81 | -0.87 | -0.99 |
|
Table 1. Adsorption energies Eads on Mn1N4/TM2 (Eads in eV)
| 3d | Sc | Ti | V | Cr | Mn | Fe | Co | Ni |
|---|
| Eads(TM2) N2 end-on
| -0.21 | -0.75 | -0.26 | -0.52 | -0.94 | -1.06 | -0.88 | -0.53 | | Eads(TM2) N2 side-on
| -0.21 | -0.37 | -0.35 | -0.41 | -0.59 | -0.54 | -0.25 | -0.56 | | Eads(TM2) H
| 0.93 | 0.33 | 0.27 | -0.02 | -0.14 | -0.25 | -0.11 | -0.37 | | 4d | Y | Zr | Nb | Mo | Tc | Ru | Rh | Pd | | Eads(TM2) N2 end-on
| -0.14 | -0.22 | -0.20 | -0.62 | -0.88 | -0.96 | -0.76 | -0.49 | | Eads(TM2) N2 side-on
| 0.22 | -0.20 | -0.20 | 0.01 | -0.58 | -0.41 | -0.27 | 0.01 | | Eads(TM2) H
| 0.90 | 0.21 | 0.17 | -0.23 | -0.12 | -0.12 | -0.09 | -0.33 | | 5d | Lu | Hf | Ta | W | Re | Os | Ir | Pt | | Eads(TM2) N2 end-on
| -0.20 | -0.31 | -0.64 | -0.91 | -1.15 | -1.27 | -1.09 | -0.26 | | Eads(TM2) N2 side-on
| -0.20 | 0.07 | -0.49 | -0.77 | -0.94 | -0.68 | -0.48 | 0.22 | | Eads(TM2) H
| 0.77 | 0.01 | -0.73 | -0.84 | -0.68 | -0.78 | -0.72 | -0.99 |
|
Table 2. Adsorption energies Eads on Fe1N4/TM2 (Eads in eV)
| 3d | Sc | Ti | V | Cr | Mn | Fe | Co | Ni |
|---|
| Eads(TM2) N2 end-on
| -0.21 | -0.37 | -0.68 | -0.84 | -1.01 | -1.05 | -0.85 | -0.46 | | Eads(TM2) N2 side-on
| -0.20 | -0.37 | -0.29 | -0.51 | -0.64 | -0.53 | -0.26 | -0.56 | | Eads(TM2) H
| 1.02 | 0.37 | -0.09 | -0.08 | -0.36 | -0.13 | -0.07 | -0.28 | | 4d | Y | Zr | Nb | Mo | Tc | Ru | Rh | Pd | | Eads(TM2) N2 end-on
| -0.12 | -0.19 | -0.44 | -0.61 | -0.82 | -0.93 | -0.75 | -0.48 | | Eads(TM2) N2 side-on
| -0.13 | -0.20 | -0.03 | -0.29 | -0.57 | -0.42 | -0.25 | -0.48 | | Eads(TM2) H
| 1.03 | 0.42 | -0.12 | -0.22 | -0.07 | -0.07 | -0.03 | -0.26 | | 5d | Lu | Hf | Ta | W | Re | Os | Ir | Pt | | Eads(TM2) N2 end-on
| -0.20 | -0.29 | -0.62 | -0.86 | -1.08 | -1.23 | -1.07 | -0.49 | | Eads(TM2) N2 side-on
| -0.21 | -0.29 | -0.28 | -0.63 | -0.89 | -0.67 | -0.46 | -0.48 | | Eads(TM2) H
| 0.82 | 0.23 | -0.50 | -0.75 | -0.63 | -0.72 | -0.69 | -0.87 |
|
Table 3. Adsorption energies Eads on Co1N4/TM2 (Eads in eV)
| 3d | Sc | Ti | V | Cr | Mn | Fe | Co | Ni |
|---|
| Eads(TM2) N2 end-on
| -0.21 | -0.41 | -0.72 | -0.91 | -1.04 | -1.07 | -0.79 | -0.58 | | Eads(TM2) N2 side-on
| -0.19 | 0.02 | -0.41 | -0.63 | -0.66 | -0.50 | / | -0.58 | | Eads(TM2) H
| 0.97 | 0.19 | -0.14 | -0.40 | -0.23 | -0.22 | -0.18 | -0.27 | | 4d | Y | Zr | Nb | Mo | Tc | Ru | Rh | Pd | | Eads(TM2) N2 end-on
| -0.12 | -0.24 | -0.51 | -0.70 | -0.91 | -0.98 | -0.73 | -0.48 | | Eads(TM2) N2 side-on
| -0.13 | -0.20 | -0.23 | -0.63 | -0.61 | -0.44 | -0.21 | -0.48 | | Eads(TM2) H
| 0.97 | 0.15 | -0.33 | -0.22 | -0.12 | -0.13 | -0.16 | -0.25 | | 5d | Lu | Hf | Ta | W | Re | Os | Ir | Pt | | Eads(TM2) N2 end-on
| -0.20 | -0.33 | -0.74 | -0.98 | -1.17 | -1.30 | -1.06 | -0.65 | | Eads(TM2) N2 side-on
| -0.20 | 0.06 | -0.49 | -0.94 | -0.95 | -0.68 | -0.41 | -0.65 | | Eads(TM2) H
| 0.85 | 0.02 | -0.71 | -0.66 | -0.69 | -0.82 | -0.81 | -0.93 |
|
Table 4. Adsorption energies Eads on Ni1N4/TM2 (Eads in eV)
| System | Mechanisms | N2 adsorption
| R1 | R2 | R3 | R4 | R5 | R6 | NH3 desorption
|
|---|
| NiN4/Cr
| Distal | -0.41 | 0.98 | -0.28 | 0.17 | -1.08 | -1.09 | -0.23 | 1.04 | | Alternating | -0.41 | 0.98 | 0.05 | -0.31 | -0.25 | -1.29 | -0.71 | 1.04 | | Enzymatic | -0.10 | 0.57 | 0.16 | -0.56 | -0.12 | -1.51 | -0.38 | 1.04 | | NiN4/Mo
| Distal | -0.27 | 0.92 | -0.08 | -0.22 | -1.14 | -0.71 | -0.20 | 1.04 | | Alternating | -0.27 | 0.92 | 0.16 | -0.56 | 0.06 | -1.52 | -0.49 | 1.04 | | Enzymatic | -0.11 | 0.60 | 0.18 | -0.89 | 0.50 | -1.54 | -0.44 | 1.04 | | NiN4/Ta
| Distal | -0.18 | 0.69 | -0.37 | -0.06 | -1.22 | -1.02 | 0.22 | 1.04 | | Alternating | -0.18 | 0.69 | 0.05 | -0.88 | 0.11 | -1.78 | 0.05 | 1.04 | | Enzymatic | 0.04 | 0.11 | -0.23 | -0.70 | 0.58 | -1.70 | -0.04 | 1.04 |
|
Table 5. Free energy change ΔG (ΔG in eV), Ri stands for the ith protonation step
| Distal | Alternating | Enzymatic |
|---|
| RDS | ΔGmax | RDS | ΔGmax | RDS | ΔGmax | | Cr | *N2+H→*NNH
| 1.03 | *N2+H→*NNH
| 1.03 | *N*N+H→*N*NH | 0.66 | | NiN4/Cr
| *N2+H→*NNH
| 0.98 | *N2+H→*NNH
| 0.98 | *N*N+H→*N*NH | 0.57 | | Mo | *N2+H→*NNH
| 1.27 | *N2+H→*NNH
| 1.27 | *N*N+H→*N*NH | 0.43 | | NiN4/Mo
| *N2+H→*NNH
| 0.92 | *N2+H→*NNH
| 0.92 | *N*N+H→*N*NH | 0.60 | | Ta | *NNH2+H→*N
| 0.72 | *N2+H→*NNH
| 0.66 | *NH*NH2+H→*NH2*NH2 | 0.49 | | NiN4/Ta
| *N2+H→*NNH
| 0.69 | *N2+H→*NNH
| 0.69 | *NH*NH2+H→*NH2*NH2 | 0.58 |
|
Table 6. Potential determining step and its free energy change ΔGmax(ΔGmax in eV)