Yiying Zhang, Yan Cao, Yuxiang Chen, Xiangwei Mu. Quality Optimization of Laser-Cutted Ni-Based Alloys Based on Genetic Algorithm[J]. Laser & Optoelectronics Progress, 2018, 55(11): 111404
Search by keywords or author
- Laser & Optoelectronics Progress
- Vol. 55, Issue 11, 111404 (2018)
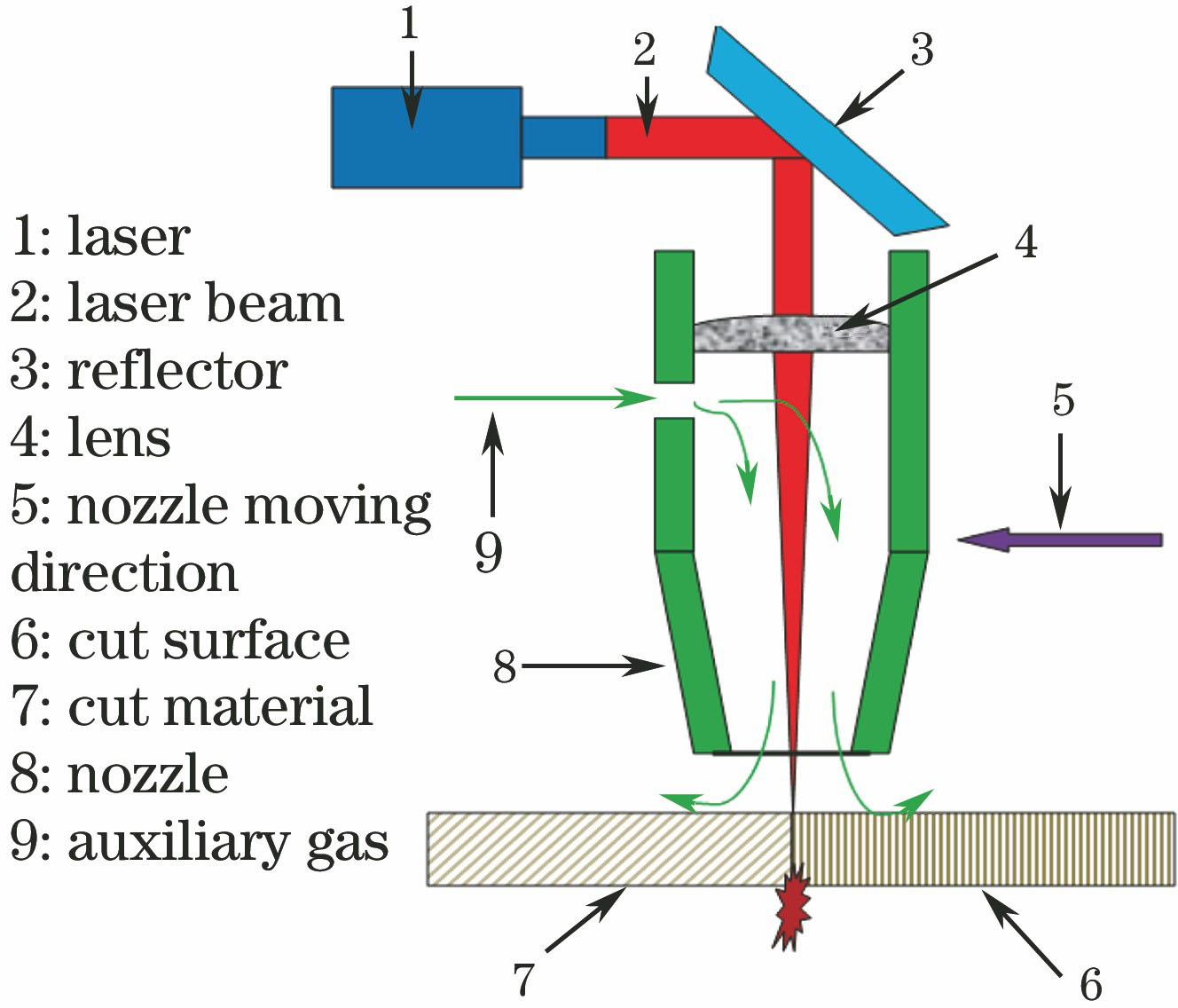
Fig. 1. Schematic of laser-cutting process
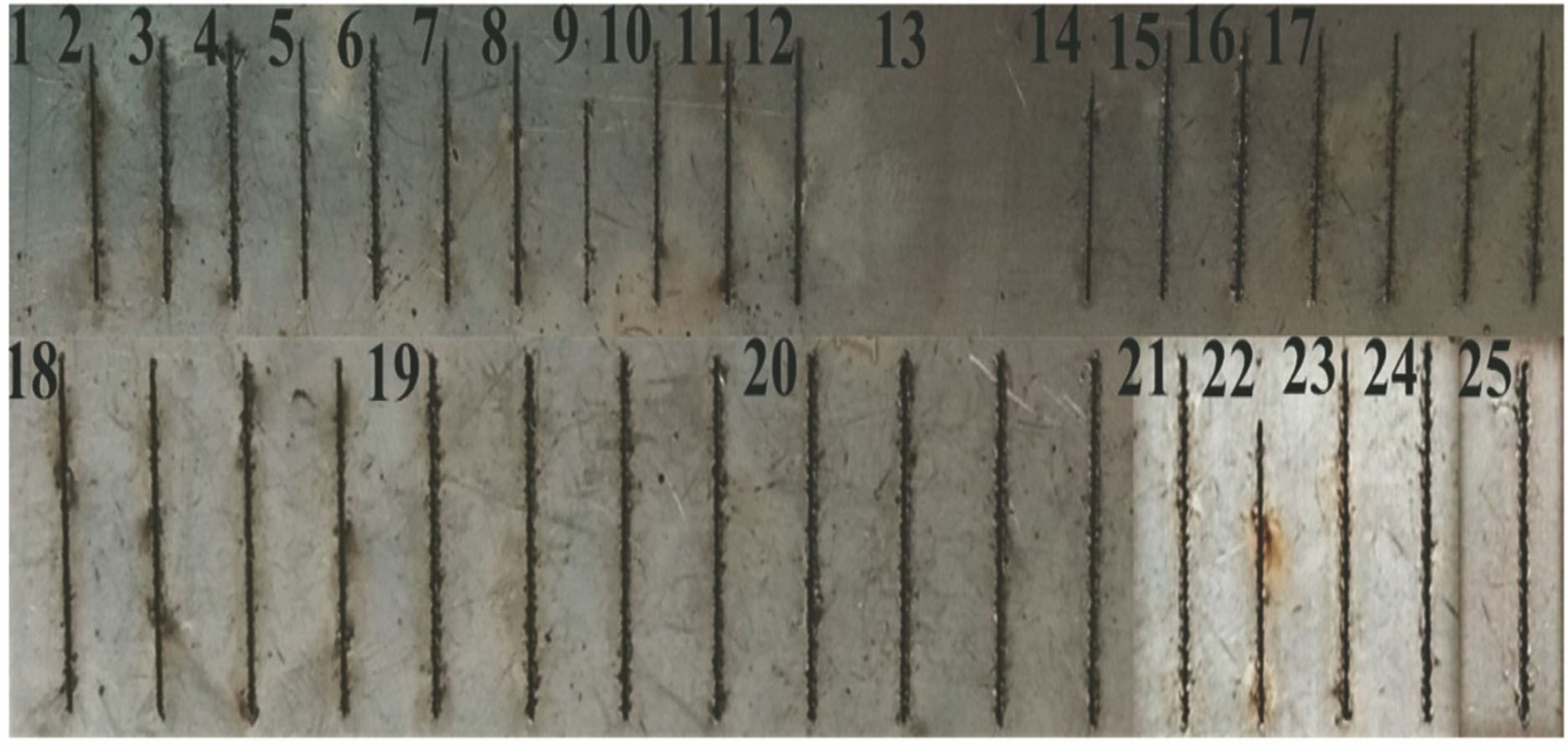

Fig. 2. Exp erimental macrograph
Fig. 3. Mark graph of metallographic data of samples. (a) No. 2; (b) No. 3; (c) No. 4; (d) No. 5; (e) No. 6; (f) No. 7; (g) No. 8; (h) No. 9; (i) No. 10; (j) No. 11; (k) No. 12; (l) No. 14; (m) No. 15; (n) No. 16; (o) No. 17; (p) No. 18; (q) No. 19; (r) No. 20; (s) No. 21; (t) No. 22; (u) No. 23; (v) No. 24; (w) No. 25
Fig. 4. Four-time sampling figures of samples from No. 17 to No. 20. (a) No. 17, first sampling; (b) No. 17, second sampling; (c) No. 17, third sampling; (d) No. 17, fourth sampling; (e) No. 18, first sampling; (f) No. 18, second sampling; (g) No. 18, third sampling; (h) No. 18, fourth sampling; (i) No. 19, first sampling; (j) No. 19, second c sampling; (k) No. 19, third sampling; (l) No. 19, fourth sampling; (m) No. 20, first sampling; (n) No. 20, second sampling; (o) No. 20, third sampling; (p) No. 20
Fig. 5. Three-layer BP network
Fig. 6. Resutls predicted by BP neural network. (a) Comparison between predicted value and expected value; (b) error; (c) percentage of error
Fig. 7. Flow chart of algorithm
Fig. 8. Fitness value curve
Fig. 9. Experimental diagram for test. (a) Positive macrograph; (b) back macrograph; (c) metallographic micrograph
|
Table 1. Chemical compositions of GH3128 (mass fraction,%)
|
Table 2. Factor levels
|
Table 3. Comprehensive scores of samples
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Table 4. Sample errors
|
Table 5. Hidden layer node errors

Set citation alerts for the article
Please enter your email address



