Ronghao Li, Yinan Chen, Xiaozheng Gan, Qing Zhang, Pei Wang. Tree-Skeleton Generation Method by Thinning Voxels of Point Cloud[J]. Laser & Optoelectronics Progress, 2019, 56(19): 192802
Search by keywords or author
- Laser & Optoelectronics Progress
- Vol. 56, Issue 19, 192802 (2019)
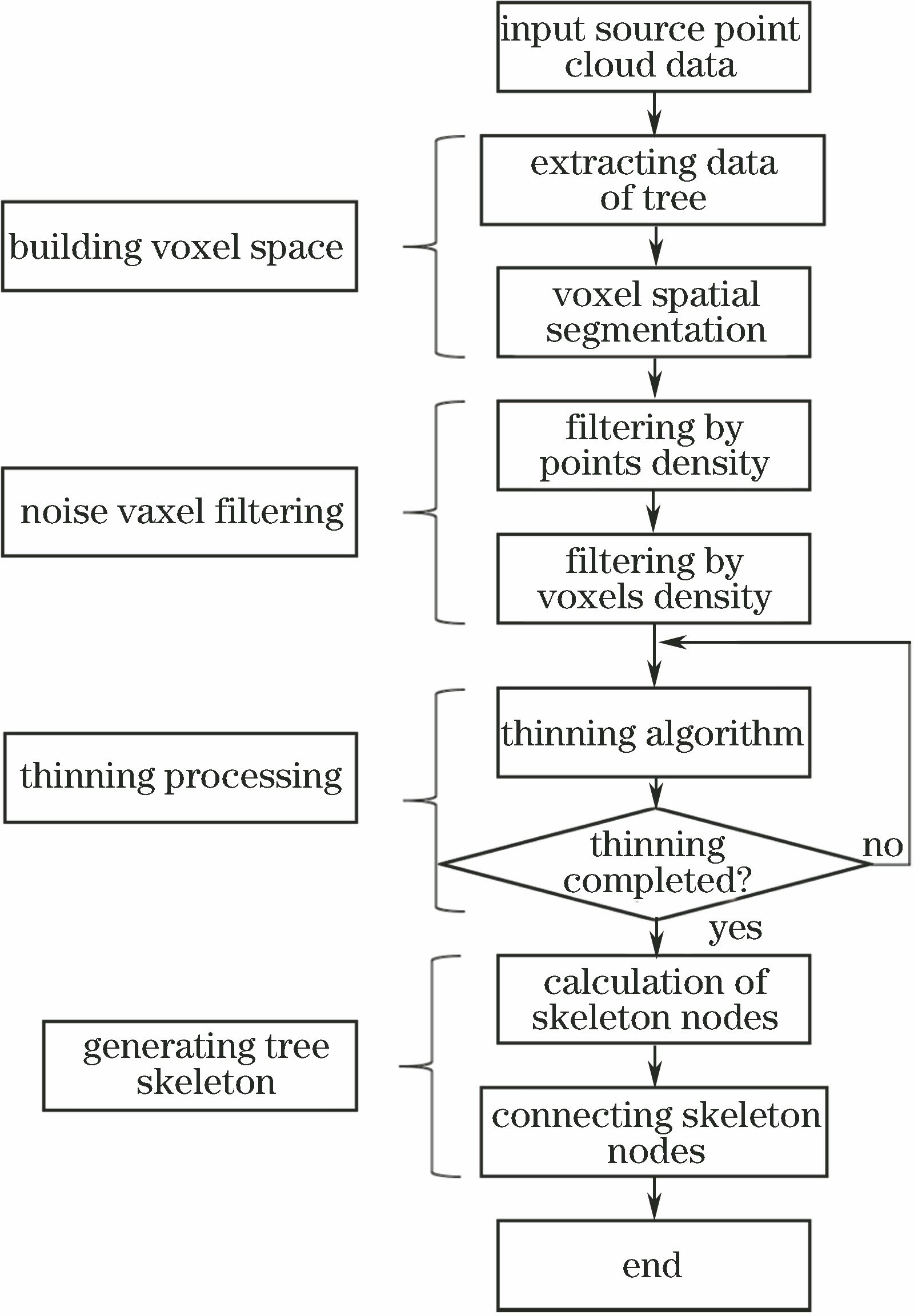
Fig. 1. Flow chart of proposed method
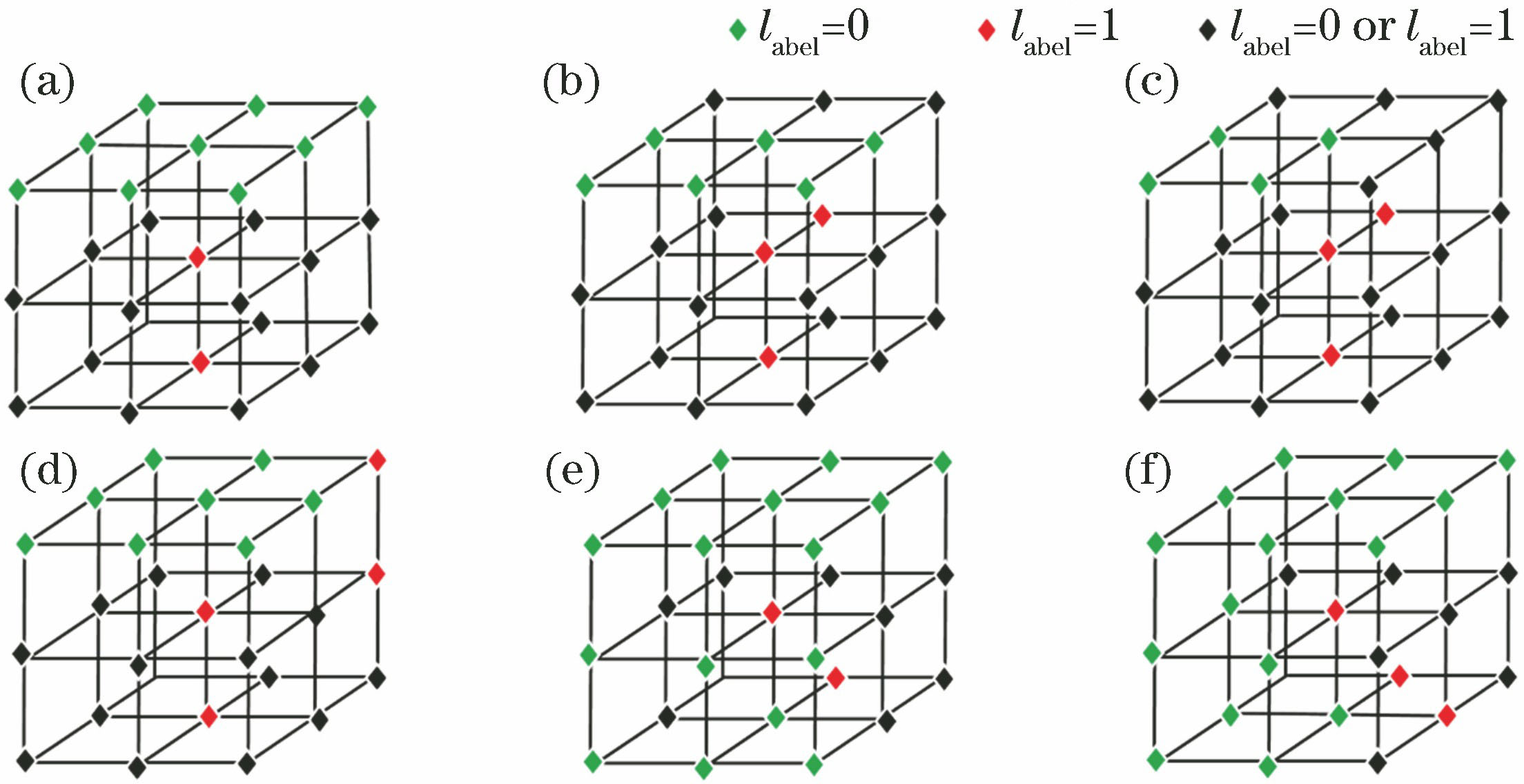
Fig. 2. Six delete templates in upper direction
Fig. 3. Flow chart of thinning algorithm
Fig. 4. Connection of skeleton nodes
Fig. 5. Generated skeletons of Ginkgo tree and Amygdalus triloba f. multiplex tree. (a) Picture of Ginkgo on left and picture of Amygdalus triloba f. multiplex on right; (b) point cloud data of Ginkgo on left and point cloud data of Amygdalus triloba f. multiplex on right; (c) generated skeleton of Ginkgo on left and generated skeleton of Amygdalus triloba f. multiplex on right
Fig. 6. Skeletons of Ginkgo tree generated by point clouds at various angular resolutions. (a) 0.02°; (b) 0.05°; (c) 0.10°
Fig. 7. Generated skeletons of Amygdalus triloba f. multiplex tree with different partition specifications. (a) (40, 40, 40); (b) (60, 60, 60); (c) (80, 80, 80); (d) (100, 100, 100)
Fig. 8. Generated skeletons of Amygdalus triloba f. multiplex tree with different filtering parameters. (a) (0,4); (b) (0,2); (c) (10,0); (d) (20,0)
Fig. 9. Tree skeletons of Ginkgo tree and Amygdalus triloba f. multiplex tree generated by GSA and IVTA methods. (a) Skeleton of Ginkgo tree generated by GSA method; (b) skeleton of Amygdalus triloba f. multiplex tree generated by GSA method; (c) skeleton of Ginkgo tree generated by IVTA method; (d) skeleton of Amygdalus triloba f. multiplex tree generated by IVTA method
|
Table 1. Acquisition information of point cloud data
|
Table 2. Skeleton information of Ginkgo tree with different angular resolutions
|
Table 3. Skeleton information of Amygdalus triloba f. multiplex tree with different partition specifications
|
Table 4. Skeleton information of Amygdalus triloba f. multiplex tree with different filtering parameters
|
Table 5. Tree skeleton informations generated by GSA and IVTA methods

Set citation alerts for the article
Please enter your email address



