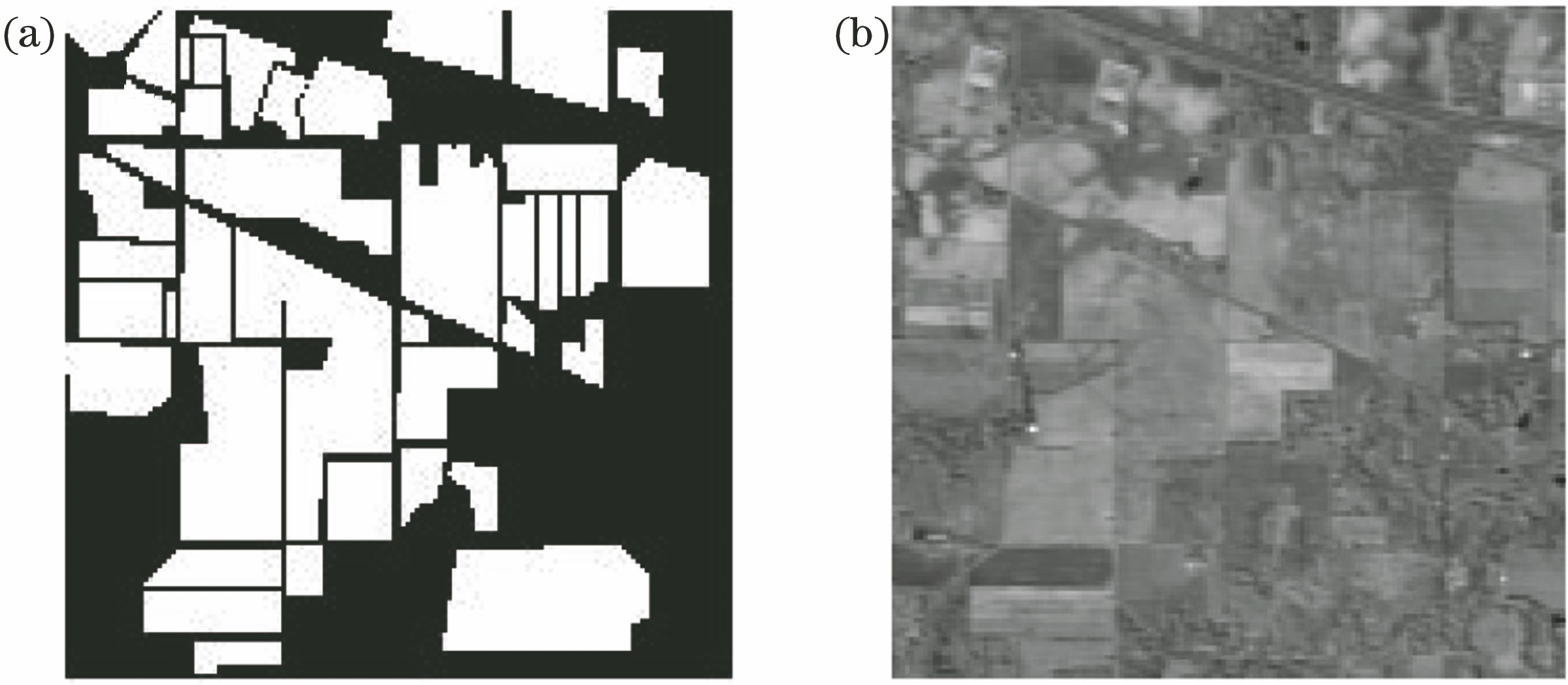
Fig. 1. Results of different feature extraction methods. (a) LBP; (b) Gabor
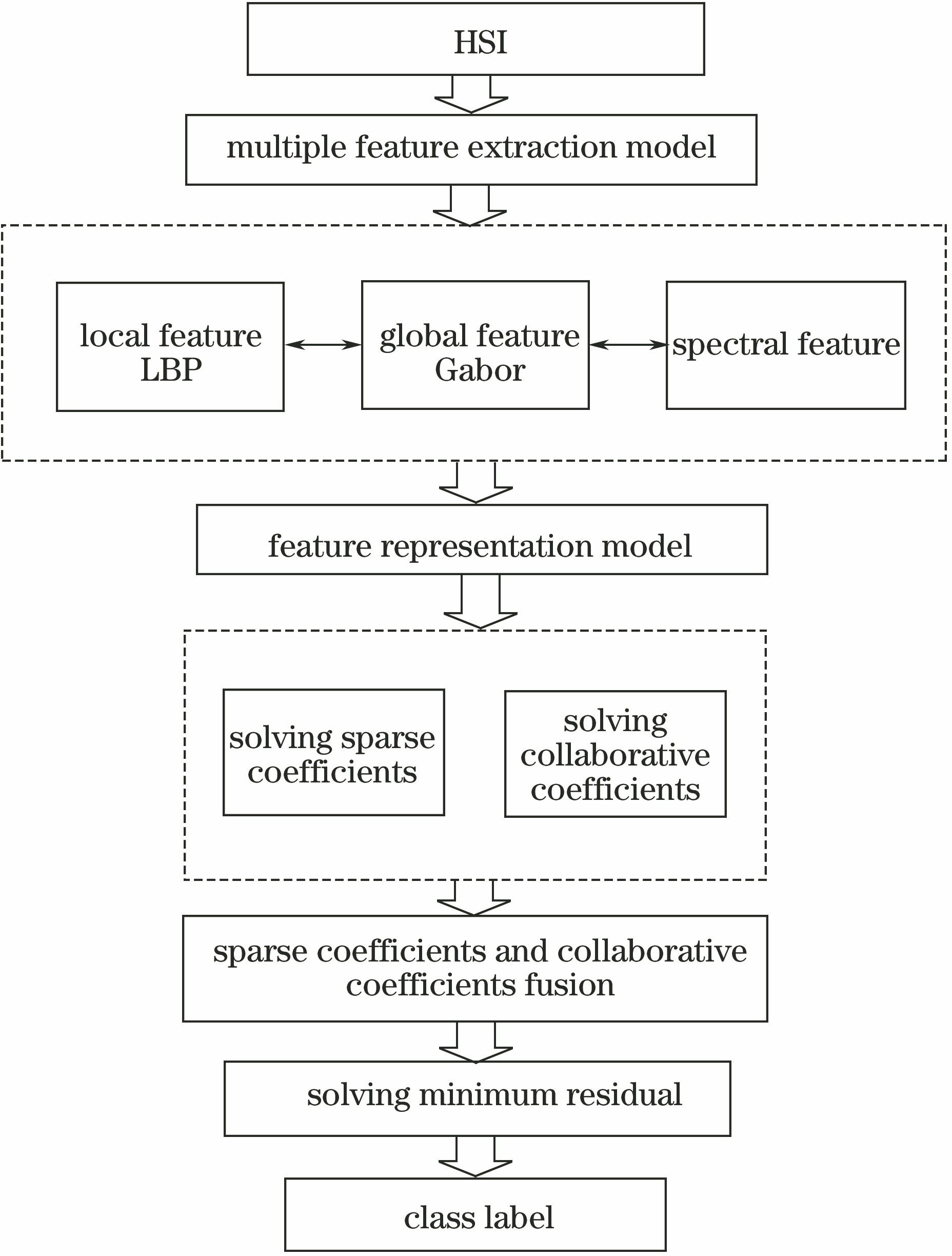
Fig. 2. Flow chart of MFISR
Fig. 3. Parameter adjustment and accuracy change
Fig. 4. Images of Salinas. (a) Pseudo-color image; (b) ground truth image
Fig. 5. Classification maps of different methods on Salinas dataset. (a) SVM; (b) SRC_Spectral; (c) SRC_Gabor; (d) SRC_LBP; (e) CRC_Spectral; (f) CRC_Gabor; (g) CRC_LBP; (h) MFISR
Fig. 6. Image of Indian Pines. (a) Pseudo-color image; (b) ground truth image
Fig. 7. Classification maps of different methods on Indian Pines dataset. (a) SVM; (b) SRC_Spectral; (c) SRC_Gabor; (d) SRC_LBP; (e) CRC_Spectral; (f) CRC_Gabor; (g) CRC_LBP; (h) MFISR
| No. | Class | Training | Test |
|---|
| 1 | Brocoli_green_weeds_1 | 41 | 1968 | | 2 | Brocoli_green_weeds_2 | 75 | 3651 | | 3 | Fallow | 40 | 1936 | | 4 | Fallow_rough_plow | 28 | 1366 | | 5 | Fallow_smooth | 54 | 2624 | | 6 | Stubble | 80 | 3879 | | 7 | Celery | 72 | 3507 | | 8 | Grapes_untrained | 226 | 11045 | | 9 | Soil_vinyard_develop | 125 | 6078 | | 10 | Corn_senesced_green_weeds | 66 | 3212 | | 11 | Lettuce_romaine_4wk | 22 | 1046 | | 12 | Lettuce_romaine_5wk | 39 | 1888 | | 13 | Lettuce_romaine_6wk | 19 | 897 | | 14 | Lettuce_romaine_7wk | 22 | 1048 | | 15 | Vinyard_untrained | 146 | 7122 | | 16 | Vinyard_vertical_trellis | 37 | 1770 | | Total | 1092 | 53037 |
|
Table 1. Training and test sample distributions of different class labels in Salinas dataset
| No. | SVM | SRC_Spectral | SRC_Gabor | SRC_LBP | CRC_Spectral | CRC_Gabor | CRD_LBP | MFISR (Spectral Value+Gabor+LBP) |
|---|
| 1 | 0.9563 | 0.8301 | 1 | 1 | 0.9918 | 0.9969 | 1 | 1 | | 2 | 0.9726 | 0.9945 | 0.9978 | 0.9997 | 0.9703 | 0.9817 | 0.9989 | 1 | | 3 | 0.8657 | 0.9770 | 0.9847 | 0.9938 | 0.9890 | 1 | 0.9954 | 1 | | 4 | 0.9810 | 0.9941 | 0.9770 | 0.9020 | 0 | 0 | 0.8713 | 0.9109 | | 5 | 0.9501 | 0.9926 | 0.9908 | 0.9767 | 0.5407 | 0.5306 | 0.9778 | 0.9519 | | 6 | 0.9773 | 0.9943 | 0.9985 | 0.9922 | 0.9852 | 0.9888 | 0.9891 | 0.9809 | | 7 | 0.9855 | 0.9864 | 0.9909 | 0.9923 | 0.9211 | 0.9372 | 0.9872 | 0.9991 | | 8 | 0.8050 | 0.7934 | 0.8096 | 0.9995 | 0.6332 | 0.6315 | 0.9954 | 0.9968 | | 9 | 0.9595 | 0.9932 | 0.9869 | 0.9975 | 0.7095 | 0.6796 | 0.9984 | 1 | | 10 | 0.7883 | 0.9535 | 0.9478 | 0.9900 | 0.8268 | 0.8792 | 0.9872 | 0.9981 | | 11 | 0.8203 | 0.9894 | 0.9311 | 0.9715 | 0.9419 | 0.9509 | 0.9581 | 0.9839 | | 12 | 0.9529 | 0.9661 | 0.9697 | 0.9700 | 0.9655 | 0.8923 | 0.9721 | 0.9951 | | 13 | 0.9588 | 0.9601 | 0.9310 | 0.9404 | 0 | 0 | 0.9100 | 0.8967 | | 14 | 0.8664 | 0.9632 | 0.9615 | 0.9561 | 0.5205 | 0.5219 | 0.9324 | 0.9538 | | 15 | 0.4985 | 0.7433 | 0.7863 | 0.9960 | 0.7073 | 0.7551 | 0.9947 | 0.9942 | | 16 | 0.8254 | 0.9886 | 0.9948 | 1 | 0.9994 | 1 | 1 | 1 | | OA | 0.8462 | 0.9068 | 0.9206 | 0.9898 | 0.7532 | 0.7508 | 0.9860 | 0.9890 | | AA | 0.8852 | 0.9450 | 0.9537 | 0.9799 | 0.7314 | 0.7341 | 0.9730 | 0.9788 |
|
Table 2. Classification accuracies of different methods on Salinas dataset
| No. | Class | Training | Test |
|---|
| 1 | Alfalfa | 5 | 41 | | 2 | Corn-notill | 143 | 1285 | | 3 | Corn-mintill | 83 | 747 | | 4 | Corn | 24 | 213 | | 5 | Grass-pasture | 49 | 434 | | 6 | Grass-trees | 73 | 657 | | 7 | Grass-pasture-mowed | 3 | 25 | | 8 | Hay-windrowed | 48 | 430 | | 9 | Oats | 2 | 18 | | 10 | Soybean-notill | 97 | 875 | | 11 | Soybean-mintill | 246 | 2209 | | 12 | Soybean-clean | 60 | 533 | | 13 | Wheat | 21 | 184 | | 14 | Woods | 127 | 1138 | | 15 | Buildings-grass-trees-drives | 39 | 347 | | 16 | Stone-steel-towers | 10 | 83 | | Total | 1030 | 9219 |
|
Table 3. Training and test sample distributions of different class labels in Indian Pines dataset
| No. | SVM | SRC_Spectral | SRC_Gabor | SRC_LBP | CRC_Spectral | CRC_Gabor | CRC_LBP | MFISR (Spectral Value+Gabor+LBP) |
|---|
| 1 | 0.7917 | 0.8947 | 0.9500 | 1 | 0 | 0 | 1 | 1 | | 2 | 0.8070 | 0.8328 | 0.9030 | 0.9750 | 0.6468 | 0.6686 | 0.9641 | 0.9793 | | 3 | 0.6720 | 0.8048 | 0.8643 | 0.9986 | 0.6352 | 0.7774 | 0.9901 | 0.9853 | | 4 | 0.6381 | 0.6376 | 0.7828 | 0.9130 | 0 | 0 | 0.8787 | 1 | | 5 | 0.8993 | 0.9486 | 0.9769 | 0.9824 | 0.9503 | 0.9932 | 0.9889 | 0.9823 | | 6 | 0.9598 | 0.8997 | 0.9395 | 0.9985 | 0.7344 | 0.7616 | 0.9970 | 0.9868 | | 7 | 0.8696 | 1 | 1 | 1 | 0 | 0 | 1 | 1 | | 8 | 0.9705 | 0.9500 | 0.9731 | 1 | 0.8318 | 0.8286 | 1 | 1 | | 9 | 0.4444 | 0.8333 | 0.9375 | 0.8500 | 0 | 0 | 0.9412 | 1 | | 10 | 0.6693 | 0.8080 | 0.8801 | 1 | 0.7727 | 0.7857 | 0.9951 | 0.9783 | | 11 | 0.7384 | 0.8366 | 0.9041 | 0.9955 | 0.5081 | 0.5436 | 0.9941 | 0.9933 | | 12 | 0.7572 | 0.8505 | 0.9245 | 0.9768 | 0.8480 | 0.8532 | 0.9572 | 0.9835 | | 13 | 0.9842 | 0.9643 | 0.9947 | 1 | 0.9895 | 1 | 1 | 1 | | 14 | 0.9296 | 0.9269 | 0.9339 | 0.9915 | 0.8471 | 0.8691 | 0.9906 | 0.9890 | | 15 | 0.6257 | 0.8354 | 0.8352 | 0.9688 | 0.9012 | 0.9524 | 0.9660 | 1 | | 16 | 0.8824 | 0.9551 | 1 | 0.9412 | 0.9048 | 0.9710 | 0.9195 | 0.9753 | | OA | 0.7957 | 0.8588 | 0.9115 | 0.9877 | 0.6593 | 0.6955 | 0.9825 | 0.9879 | | AA | 0.7899 | 0.7997 | 0.8906 | 0.9652 | 0.7975 | 0.6253 | 0.9739 | 0.9908 |
|
Table 4. Classification accuracies of different methods on Indian Pines dataset