zhenzhen Wan, Shaoyong Zhang, Fang Liu, Ning Shi, Shuai Han, Chunxue Li. Computer-Aided Evaluation of Ki-67 on Pathological Sections of Appendiceal Neuroendocrine Neoplasms[J]. Laser & Optoelectronics Progress, 2021, 58(24): 2417001
Search by keywords or author
- Laser & Optoelectronics Progress
- Vol. 58, Issue 24, 2417001 (2021)
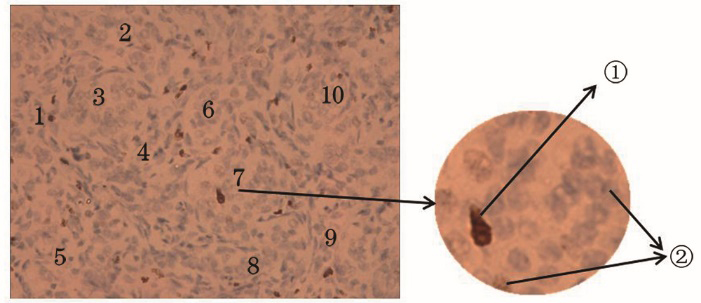
Fig. 1. Pathological section of ANENs
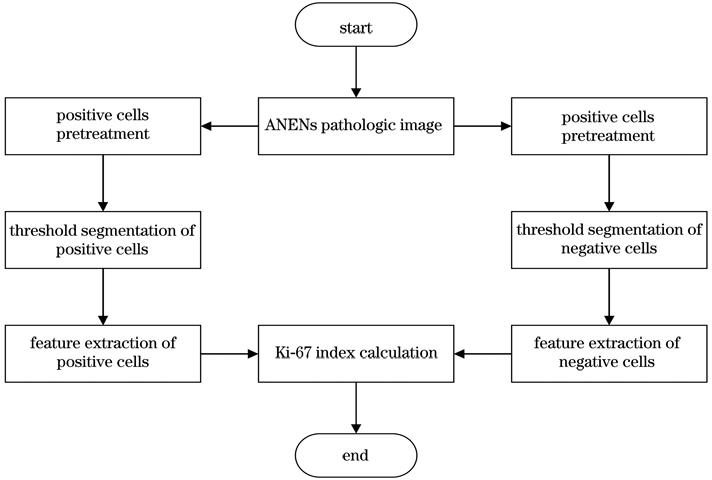
Fig. 2. Overall design flow chart of the system scheme
Fig. 3. Image of ANENs pathological section (left) and color image of YUV (right)
Fig. 4. YUV spatial threshold segmentation images. (a) Threshold segmentation image of positive cell; (b) threshold segmentation image of negative cells
Fig. 5. Images processed by mathematical morphology. (a) Positive cell; (b) negative cells
Fig. 6. One-dimensional minimum highlight process. (a) One-dimensional data; (b) minimum highlight results
Fig. 7. Positive cells processing images obtained by improved watershed algorithm. (a) Image of ANENs pathological section; (b) distance transform image; (c) image by forced minimum technique processing; (d) image processed by improved watershed algorithm; (e) binary segmentation image; (f) color marker segmentation image
Fig. 8. Negative cells processing images obtained by improved watershed algorithm. (a) Image of ANENs pathological section; (b) distance transform images; (c) image by forced minimum technique processing; (d) image processed by improved watershed algorithm; (e) binary segmentation image; (f) color marker segmentation image
Fig. 9. Negative cells segmentation effect diagrams before and after algorithm improvement. (a)--(c) Traditional watershed algorithm; (d)--(f) improved watershed algorithm
Fig. 10. Over-segmentation rate data of negative cells
Fig. 11. Data analysis of Ki-67 index
Fig. 12. Comparison of time consumption on image processing
| ||||||||||||||||||||||||||||||||||||||||||||||||
Table 1. Experimental data of positive cells in ANENs pathological section images obtained by improved watershed algorithm
|
Table 2. Experimental data of negative cells in ANENs pathological section images
|
Table 3. Experimental data of diagnostic indicators of ANENs pathological sections

Set citation alerts for the article
Please enter your email address



