- Opto-Electronic Advances
- Vol. 6, Issue 6, 230083 (2023)
Abstract
TPM is a non-invasive imaging technique that combines digital holography and tomography to provide three-dimensional (3D) images of transparent samples based on their refractive index (RI) contrast
Overall, this QPI-based TPM approach has the potential to promote greater accessibility in lipid droplet research and enable the development of more advanced LD-assisted biomedical tools.
Furthermore, such comprehensive characterization of LDs may assist in their function as endogenous microlenses
![]()
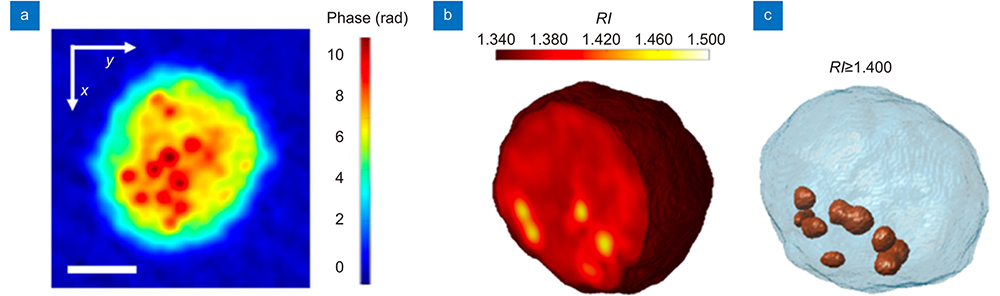
Figure 1.(
In their recently published paper in Opto-Electronic Advances, Pietro Ferraro and his colleagues report on a new high-throughput tomographic phase instrument that precisely quantifies intracellular lipid droplets (LDs)
LDs are lipid storage organelles found in most cell types and play an active role in critical biological processes, including energy metabolism, membrane homeostasis, and biomolecular signaling
In this recent work by Ferraro et al., TPM was applied for visualizing and characterizing LDs in stain-free suspended cells flowing through a microfluidic channel. The devised in-flow TPM system was proven reliable by reconstructing the 3D refractive index distribution of LDs in ovarian cancer (
Prospectively, this technology establishes the groundwork for a plethora of exciting imaging opportunities with lipid droplets, especially if used in conjunction with deep learning models. For instance, the average in-flow TPM experiment involves recording holographic sequences comprising hundreds of images with 3D RI reconstructions typically requiring about 15 minutes. With fewer frames, this step can potentially be accelerated by implementing appropriate neural networks. In addition, this microfluidic system propels hundreds of cells past the sensor per minute, at 50 nL/s, and is an ideal setup for achieving substantially higher throughput with the aid of machine learning. For example, its throughput may be increased using specialized deblurring neural networks trained on data acquired at different flow rates
These measurements enabled the extraction of important parameters related to spatial correlations and three-dimensional (3D) positioning of LDs at high-volume rates. Collecting such information from numerous cells yields statistically rich data, revealing additional quantitative properties, such as the RI, equivalent radius, dry mass, sphericity, and concentric LD zones. Beyond these metrics, the 3D spatial localization of LDs inside suspended cells was also delineated, which is advantageous to understanding the interactions of LDs with other organelles. Also, this opto-fluidic TPM configuration is highly conducive to monitoring various simulated physiological scenarios, such as circulating tumor cells.
Transmission electron microscopy (TEM) and fluorescence microscopy (FM) are the two most commonly employed techniques for investigating LDs
To address these limitations, label-free methods, such as Tomographic Phase Microscopy (TPM), have been increasingly adopted as a viable imaging alternative, especially for live cells. Moreover, coupled with deep learning-based computational postprocessing, label-free modalities have been successfully used to circumvent many traditional chemical preparations of biological specimens and avoid the related cost, labor, duration, sample damage and inconsistencies of such procedures
References
[1] D Pirone, D Sirico, L Miccio, V Bianco, M Mugnano et al. 3D imaging lipidometry in single cell by in-flow holographic tomography. Opto-Electronic Adv, 6, 220048(2023).
[2] JA Olzmann, P Carvalho. Dynamics and functions of lipid droplets. Nat Rev Mol Cell Biol, 20, 137-155(2019).
[3] Y Imai, RS Cousins, SM Liu, BM Phelps, JA Promes. Connecting pancreatic islet lipid metabolism with insulin secretion and the development of type 2 diabetes. Ann N Y Acad Sci, 1461, 53-72(2020).
[4] ALS Cruz, EDA Barreto, NPB Fazolini, JPB Viola, PT Bozza. Lipid droplets: platforms with multiple functions in cancer hallmarks. Cell Death Dis, 11, 105(2020).
[5] SSG Dias, VC Soares, AC Ferreira, CQ Sacramento, N Fintelman-Rodrigues et al. Lipid droplets fuel SARS-CoV-2 replication and production of inflammatory mediators. PLoS Pathog, 16, e1009127(2020).
[6] AS Martins, IC Martins, NC Santos. Methods for lipid droplet biophysical characterization in Flaviviridae infections. Front Microbiol, 9, 1951(2018).
[7] BJ Bai, XL Yang, YZ Li, YJ Zhang, N Pillar et al. Deep learning-enabled virtual histological staining of biological samples. Light Sci Appl, 12, 57(2023).
[8] YJ Rivenson, XL Yang, YZ Li, YJ Zhang, N Pillar et al. Virtual histological staining of unlabelled tissue-autofluorescence images via deep learning. Nat Biomed Eng, 3, 466-477(2019).
[9] Y Rivenson, TR Liu, ZS Wei, YB Zhang, K De Haan et al. PhaseStain: the digital staining of label-free quantitative phase microscopy images using deep learning. Light Sci Appl, 8, 23(2019).
[10] W Choi, C Fang-Yen, K Badizadegan, S Oh, N Lue et al. Tomographic phase microscopy. Nat Methods, 4, 717-719(2007).
[11] PA Sandoz, C Tremblay, FG van der Goot, M Frechin. Image-based analysis of living mammalian cells using label-free 3D refractive index maps reveals new organelle dynamics and dry mass flux. PLoS Biol, 17, e3000553(2019).
[12] MJ Fanous, SH He, S Sengupta, K Tangella, N Sobh et al. White blood cell detection, classification and analysis using phase imaging with computational specificity (PICS). Sci Rep, 12, 20043(2022).
[13] MJ Fanous, YF Li, ME Kandel, AA Abdeen, KA Kilian et al. Effects of substrate patterning on cellular spheroid growth and dynamics measured by gradient light interference microscopy (GLIM). J Biophotonics, 12, e201900178(2019).
[14] YF Li, MJ Fanous, KA Kilian, G Popescu. Quantitative phase imaging reveals matrix stiffness-dependent growth and migration of cancer cells. Sci Rep, 9, 248(2019).
[15] SO Isikman, W Bishara, S Mavandadi, FW Yu, S Feng et al. Lens-free optical tomographic microscope with a large imaging volume on a chip. Proc Natl Acad Sci USA, 108, 7296-7301(2011).
[16] W Bishara, SO Isikman, A Ozcan. Lensfree optofluidic microscopy and tomography. Ann Biomed Eng, 40, 251-262(2012).
[17] MJ Fanous, G Popescu. GANscan: continuous scanning microscopy using deep learning deblurring. Light Sci Appl, 11, 265(2022).
[18] XX Chen, TL Wu, ZY Gong, JH Guo, XS Liu et al. Lipid droplets as endogenous intracellular microlenses. Light Sci Appl, 10, 242(2021).
Set citation alerts for the article
Please enter your email address



